CRISPR/Cas9 for Alternative Splicing Mutations: A Researcher's Guide to Modeling, Correction, and Therapeutic Applications
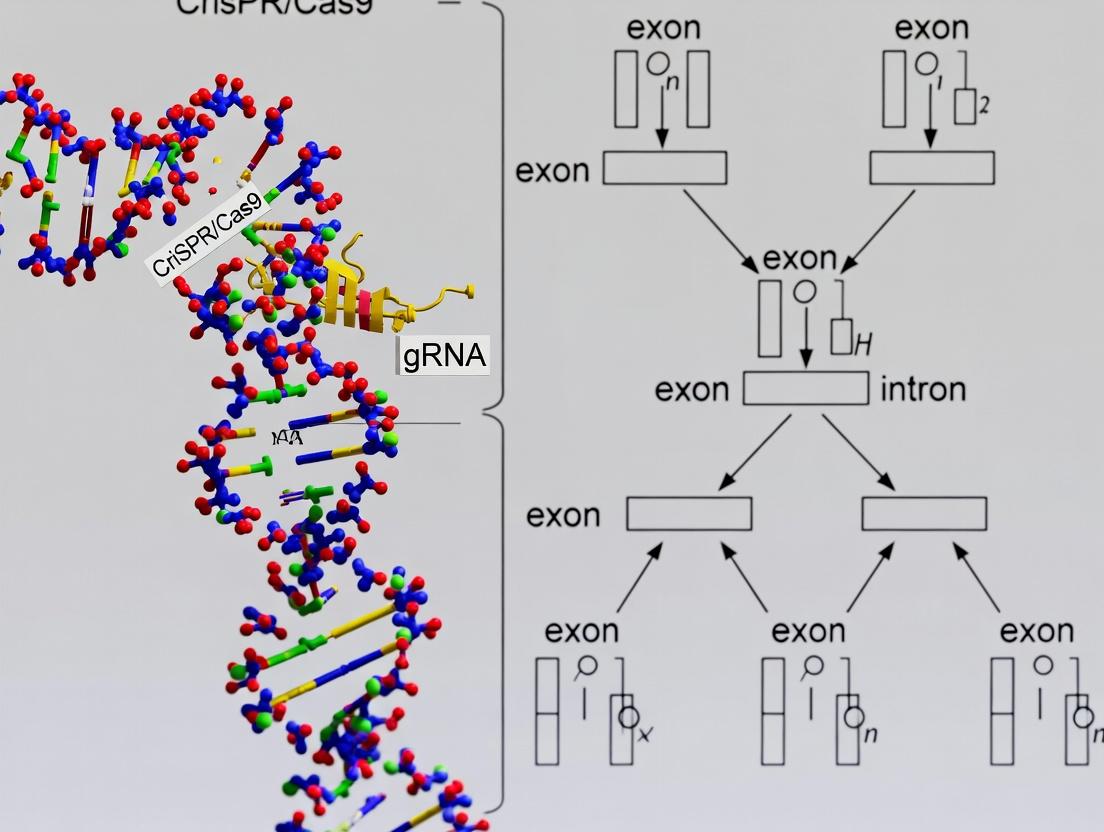
This comprehensive guide for researchers, scientists, and drug development professionals explores the pivotal role of CRISPR/Cas9 technology in studying and correcting disease-causing alternative splicing mutations.
CRISPR/Cas9 for Alternative Splicing Mutations: A Researcher's Guide to Modeling, Correction, and Therapeutic Applications
Abstract
This comprehensive guide for researchers, scientists, and drug development professionals explores the pivotal role of CRISPR/Cas9 technology in studying and correcting disease-causing alternative splicing mutations. It covers foundational knowledge on splicing mutation mechanisms, detailed methodologies for CRISPR-based modeling and correction, essential troubleshooting and optimization strategies for splicing-specific editing, and robust validation frameworks. The article synthesizes the latest research to provide a roadmap for utilizing CRISPR/Cas9 to understand splicing biology and develop novel genetic therapies, addressing both basic research and translational drug discovery needs.
Understanding the Target: The Biology and Pathology of Alternative Splicing Mutations
Alternative splicing (AS) is a fundamental post-transcriptional mechanism enabling single genes to produce multiple mRNA isoforms, dramatically expanding proteomic diversity from a limited genome. In humans, over 95% of multi-exon genes undergo AS, making it a critical regulator of development, cell differentiation, and disease.
Table 1: Quantitative Landscape of Human Alternative Splicing
| Metric | Value | Reference/Note |
|---|---|---|
| Protein-coding genes subject to AS | >95% | Wang et al., Nature Rev Genet, 2024 |
| Major AS types (SE, A5SS, A3SS, MXE, RI) | 5 | Common classification |
| Avg. isoforms per multi-exon gene | ~7 | Latest long-read sequencing data |
| Splicing-related human diseases | >15% of point mutations | From HGMD database |
| AS events dysregulated in cancer | Thousands per tumor | Pan-cancer analyses (TCGA) |
Core Mechanisms and CRISPR/Cas9 Research Context
AS is orchestrated by the spliceosome and auxiliary splicing factors (SFs) recognizing cis-regulatory elements: Exonic/Intronic Splicing Enhancers (ESEs/ISEs) and Silencers (ESSs/ISSs). In CRISPR/Cas9-based AS mutation research, the goal is to model disease or correct pathogenic splice-disrupting mutations by precisely editing these genomic elements.
Table 2: Key Cis-Elements for CRISPR/Cas9 Targeting in AS Research
| Element | Sequence Motif | Typical Function | CRISPR Application |
|---|---|---|---|
| 5' Splice Site (5'ss) | AG|GURAGU | Exon recognition | Correct donor site mutations |
| 3' Splice Site (3'ss) | YAG|R | Exon definition | Correct acceptor site mutations |
| Branch Point (BP) | CURAY | Lariat formation | Model BP mutation diseases |
| Exonic Splicing Enhancer (ESE) | e.g., (GAR)n | Bind SR proteins, promote inclusion | Disrupt to induce exon skipping |
| Exonic Splicing Silencer (ESS) | Variable | Bind hnRNPs, promote skipping | Delete to restore exon inclusion |
Detailed Application Notes & Protocols
Protocol 3.1: Designing CRISPR/Cas9 for Alternative Splicing Mutation Modeling
Objective: Introduce a point mutation within a cis-regulatory element (e.g., an ESE) to disrupt normal splicing patterns in a cell line.
Materials & Reagents:
- sgRNA Design Tools: CHOPCHOP, CRISPRdirect, Benchling.
- Cas9 Expression System: Plasmid expressing SpCas9 (e.g., pSpCas9(BB)-2A-Puro).
- sgRNA Cloning Vector: e.g., pU6-(BbsI)_CBh-Cas9-T2A-mCherry.
- HDR Template: Single-stranded oligodeoxynucleotide (ssODN) containing the desired point mutation and homologous arms (~60 nt each side).
- Cell Line: Relevant adherent cell line (e.g., HEK293T, Hela).
- Transfection Reagent: Lipofectamine 3000 or nucleofection kit.
- Validation Primers: PCR primers flanking the target site and spanning the alternative exon.
Procedure:
- Target Identification: Identify the cis-element (ESE/ESS) via databases like ESEfinder or RegSNP-intron. Design two sgRNAs flanking the element to excise it, or one sgRNA near the point mutation for HDR.
- sgRNA Cloning: Synthesize oligos, anneal, and ligate into the BbsI-linearized sgRNA vector. Transform, sequence-verify.
- HDR Template Design: Order an ssODN with the pathogenic point mutation (e.g., a G>A transition disrupting an ESE hexamer). Include a silent PAM-disrupting mutation if possible.
- Cell Transfection: Co-transfect cells (in a 12-well plate) with 500 ng Cas9 plasmid, 250 ng sgRNA plasmid, and 100 pmol of ssODN using Lipofectamine. Include controls (Cas9 only, sgRNA only).
- Isolation and Screening: 48-72h post-transfection, puromycin select (if applicable). Harvest genomic DNA. Perform PCR with flanking primers and Sanger sequence. Use TIDE or ICE analysis to quantify editing efficiency.
- Splicing Analysis: Isolate total RNA (TRIzol), treat with DNase I, reverse transcribe. Perform RT-PCR with primers in constitutive exons flanking the alternative exon. Resolve products on high-percentage agarose (3%) or Bioanalyzer. Quantify isoform ratios.
Protocol 3.2: Detecting Altered Splicing Isoforms Post-Editing
Objective: Quantitatively assess changes in splicing ratios after CRISPR/Cas9-mediated mutation.
Method: Reverse Transcription-PCR (RT-PCR) & Capillary Electrophoresis
- cDNA Synthesis: Use 1 µg total RNA, random hexamers, and a reverse transcriptase with high fidelity (e.g., SuperScript IV).
- Fluorescent PCR: Perform PCR with gene-specific primers (one 5'-labeled with 6-FAM). Use a low cycle number (22-28 cycles) in the linear range.
- Fragment Analysis: Dilute PCR product and run on a capillary electrophoresis sequencer (e.g., ABI 3730). Include size standard.
- Data Analysis: Use software (e.g., GeneMapper) to quantify peak areas corresponding to different isoform sizes. Calculate Percent Spliced In (PSI or Ψ) for exon inclusion: Ψ = (Inclusion peak area / (Inclusion + Exclusion peak areas)) * 100.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for CRISPR/Cas9 Splicing Research
| Item | Function & Rationale | Example Product/Catalog # |
|---|---|---|
| High-Efficiency Cas9 | Generates DSB for NHEJ/HDR. High activity is critical for hard-to-edit loci. | SpCas9 (TrueCut Cas9 Protein v2) |
| Chemically Modified sgRNA | Increases stability and editing efficiency, especially with RNP delivery. | Synthego 2.0 CRISPR sgRNA |
| HDR Enhancers | Small molecules that inhibit NHEJ or promote HDR, boosting precise editing. | Alt-R HDR Enhancer V2 (IDT) |
| Splicing-Sensitive Reporter | Rapid, quantitative readout of splicing changes from edited cells. | pSpliceExpress or minigene constructs |
| Splicing Factor Antibodies | Validate changes in splicing machinery via WB/IP after editing. | Anti-SRSF1, Anti-hnRNP A1 (Abcam) |
| Long-Read Sequencing Kit | Unambiguously characterize full-length mRNA isoforms post-editing. | Oxford Nanopore cDNA-PCR Sequencing Kit |
| RT-qPCR Assays for Isoforms | Absolute quantification of specific splice variant expression. | TaqMan assays spanning exon junctions |
Visualizations
Diagram Title: Central Dogma and Splicing with CRISPR Intervention
Diagram Title: CRISPR/Cas9 Splicing Mutation Workflow
Diagram Title: Targeting Cis-Elements with CRISPR/Cas9
Pathogenic splicing mutations, which constitute over 30% of disease-causing genetic variants, disrupt the precision of pre-mRNA processing. These mutations operate through two primary mechanistic categories: (1) disruption of cis-regulatory elements (e.g., splice donor/acceptor sites, branch points, exonic and intronic splicing enhancers/silencers), and (2) disruption or alteration of trans-acting splicing factors (e.g., SNRNPs, hnRNPs, SR proteins). Within our broader thesis on CRISPR/Cas9-mediated correction of aberrant splicing, understanding these mechanisms is critical for designing precise therapeutic interventions. The following application notes and protocols detail methodologies for dissecting these mechanisms and creating cellular models for drug screening.
Table 1: Prevalence and Impact of Major Splicing Mutation Types
| Mutation Type | Approximate % of Pathogenic Variants | Common Disease Associations | Typical Effect on Splicing |
|---|---|---|---|
| Splice Site (Donor/Acceptor) | 15-20% | Cystic Fibrosis (CFTR), Spinal Muscular Atrophy (SMN1) | Exon skipping, intron retention, cryptic site use |
| Branch Point Mutation | ~5% | Hereditary Hemochromatosis (HFE), Retinitis Pigmentosa | Intron retention, reduced splicing efficiency |
| Exonic Splicing Enhancer (ESE) Disruption | 8-12% | Familial Dysautonomia (IKBKAP), Duchenne Muscular Dystrophy (DMD) | Exon skipping |
| Exonic/Intronic Splicing Silencer (ESS/ISS) Creation | 5-10% | Tay-Sachs disease (HEXA), Neurofibromatosis Type 1 (NF1) | Exon skipping, altered isoform ratio |
| Trans-Factor Gene Mutation (e.g., SF3B1, U2AF1) | Varies by cancer | Myelodysplastic Syndromes, Chronic Lymphocytic Leukemia | Global splicing alteration, specific cassette exon changes |
Table 2: CRISPR/Cas9 Editing Outcomes for Splicing Correction (Representative Studies)
| Target Disease | Mutation Type | Correction Strategy (via HDR) | Reported Splicing Restoration Efficiency* | Reference Year |
|---|---|---|---|---|
| Cystic Fibrosis | CFTR c.3718-2477C>T (3849+10kb C>T) | Cryptic exon exclusion via donor site disruption | 40-60% WT transcript | 2022 |
| Spinal Muscular Atrophy | SMN2 exon 7 skipping (ISS) | ESE strengthening & ISS weakening | Up to 80% exon 7 inclusion | 2023 |
| Duchenne Muscular Dystrophy | DMD exon 45-55 deletion frame-shift | Multi-exon skipping via acceptor/donor disruption | ~70% targeted skipping (in vitro) | 2023 |
| Beta-Thalassemia | HBB IVS1-110 G>A | Cryptic splice site elimination | ~50% normal splicing | 2021 |
*Efficiencies are highly dependent on cell type, delivery method, and guide RNA design.
Experimental Protocols
Protocol 3.1:In SilicoIdentification and Prioritization of Pathogenic Splicing Variants
Purpose: To bioinformatically analyze genetic variants for potential splicing disruption.
- Variant Input: Compile VCF files from patient sequencing (Whole Genome or Exome).
- Splicing Effect Prediction: Process variants through a pipeline: a. SpliceAI (v1.3): Score for donor/acceptor loss/gain (threshold: >0.2 probability). b. MMSplice & SPIDEX: Quantify effect on splicing efficiency and ΔΨ (psi) score. c. ESEFinder/ RESCUE-ESE: Scan for ESE/ESS motif disruption or creation.
- Conservation & Integration: Cross-reference with PhyloP scores and clinical databases (ClinVar, gnomAD). Prioritize variants disrupting highly conserved canonical sites or creating strong cryptic sites.
Protocol 3.2: Functional Validation via Minigene Splicing Assay
Purpose: To experimentally test the impact of a candidate variant on splicing.
- Minigene Construction: Clone a genomic fragment (300-500 bp flanking the exon of interest) into an exon-trapping vector (e.g., pSpliceExpress or pET01).
- Site-Directed Mutagenesis: Introduce the patient-derived mutation into the wild-type minigene construct.
- Cell Transfection: Transfect wild-type and mutant minigenes into relevant cell lines (HEK293T, HeLa, or disease-specific iPSCs) in triplicate.
- RNA Analysis: Isolate total RNA 48h post-transfection. a. RT-PCR: Use vector-specific primers flanking the cloned insert. b. Gel Electrophoresis: Resolve PCR products on a 2-3% agarose gel. c. Quantification: Analyze band intensities to calculate Percent Spliced In (PSI) using ImageJ or similar software. Confirm products by Sanger sequencing.
Protocol 3.3: CRISPR/Cas9-Mediated Correction in Cellular Models
Purpose: To correct a pathogenic splicing mutation in patient-derived iPSCs via homology-directed repair (HDR).
- gRNA Design: Design two sgRNAs flanking the mutation using online tools (e.g., CRISPick, CHOPCHOP). Target as close as possible to the variant.
- Donor Template Design: Synthesize a single-stranded oligodeoxynucleotide (ssODN) donor template (~200 nt) containing the corrected sequence, with synonymous "blocker" mutations in the PAM site to prevent re-cutting.
- Electroporation: Co-electroporate patient iPSCs with Cas9 ribonucleoprotein (RNP) complexes (sgRNA + SpCas9 protein) and the ssODN donor using a Neon Transfection System.
- Clone Isolation & Screening: Single-cell sort into 96-well plates. Expand clones for 2-3 weeks. a. Genotyping: Perform PCR on genomic DNA and sequence to identify homozygous corrected clones. b. RT-PCR & Sequencing: Confirm restoration of normal splicing patterns from the endogenous locus. c. Off-target Analysis: Perform targeted sequencing of top 5 predicted off-target sites for validated clones.
Diagrams
Title: Mechanisms of Splicing Disruption by Cis and Trans Mutations
Title: CRISPR Workflow for Splicing Mutation Correction
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Splicing Mutation Research
| Item | Function/Application | Example Product/Supplier |
|---|---|---|
| Exon-Trapping Minigene Vectors | Functional validation of cis-acting variants via transient transfection. | pSpliceExpress (Addgene), pET01 (MoBiTec) |
| Splice-Sensitive RT-PCR Kits | Detection and quantification of alternative splicing isoforms from RNA. | OneStep RT-PCR Kit (Qiagen), SuperScript IV (Thermo) |
| CRISPR-Cas9 Ribonucleoprotein (RNP) | High-efficiency, transient editing with reduced off-target effects. | Alt-R S.p. Cas9 Nuclease V3 (IDT), TrueCut Cas9 Protein v2 (Thermo) |
| Chemically Modified ssODN Donors | HDR template for precise correction; chemical modifications enhance stability. | Ultramer DNA Oligos (IDT), CRISPR HDR Enhancer (Takara) |
| Splicing Reporter Cell Lines | High-throughput screening for modulators of specific splicing events. | Luciferase-based Splicing Reporters (SwitchGear Genomics) |
| Anti-Splicing Factor Antibodies | RIP-seq, CLIP-seq, or western blot to study trans-factor binding/expression. | Anti-SRSF2 (Abcam), Anti-hnRNP A1 (Santa Cruz) |
| Nanopore Direct RNA-Seq Kits | Long-read sequencing to capture full-length splice isoforms without artifacts. | Direct RNA Sequencing Kit (Oxford Nanopore) |
| Splicing-Targeted Antisense Oligos (ASOs) | Experimental control to modulate splicing (e.g., induce exon skipping/inclusion). | Morpholinos (Gene Tools), GapmeRs (Qiagen) |
Within the broader thesis on CRISPR/Cas9-mediated correction of alternative splicing mutations, this application note details the molecular pathology, quantitative impact, and experimental protocols for key disorders where aberrant splicing is a primary driver. Spinal Muscular Atrophy (SMA), Duchenne Muscular Dystrophy (DMD), and numerous cancers serve as paradigms for understanding and targeting splicing defects.
Pathogenic Mechanisms and Quantitative Impact
Aberrant splicing events disrupt the production of functional proteins, leading to disease. The table below summarizes the core splicing defects and their quantifiable consequences.
Table 1: Key Disorders and Aberrant Splicing Metrics
| Disorder | Gene | Aberrant Splicing Event | Functional Consequence | Prevalence/Incidence Data | Protein Loss/Abnormality |
|---|---|---|---|---|---|
| Spinal Muscular Atrophy (SMA) | SMN1 | Exon 7 skipping in SMN2 paralog due to a C-to-T transition in exon 7 (c.840C>T). | Loss of functional Survival Motor Neuron (SMN) protein. | ~1 in 10,000 live births; carrier frequency 1/40-1/60. | <10% of normal SMN protein levels in severe Type I. |
| Duchenne Muscular Dystrophy (DMD) | DMD | Exon skipping (e.g., exons 45-55, 51, 53) due to nonsense or frameshift mutations. | Out-of-frame transcripts, premature termination, loss of dystrophin. | ~1 in 3,500-5,000 male births worldwide. | Dystrophin typically <3% of normal in muscle biopsies. |
| Cancer (e.g., Chronic Lymphocytic Leukemia, CLL) | SF3B1 | Mutations in spliceosome component lead to widespread mis-splicing (e.g., MAP3K7 3'SS usage). | Genomic instability, altered cell signaling, oncogenesis. | SF3B1 mutations in ~20-30% of CLL, 15-20% of MDS. | Altered protein isoforms driving proliferation/survival. |
Detailed Experimental Protocols
Protocol 1:In VitroSplicing Assay for Splice-Switching Oligonucleotide (SSO) Validation
Objective: To validate the efficacy of antisense oligonucleotides (ASOs) designed to correct aberrant exon skipping (e.g., for SMN2 or DMD). Materials:
- Minigene construct containing the target exon with flanking intronic sequences.
- HeLa or HEK293T cells.
- Lipofectamine 3000 transfection reagent.
- Experimental and control ASOs (e.g., targeting ISS-N1 in SMN2).
- TRIzol reagent for RNA isolation.
- RT-PCR kit with fluorescent dyes (e.g., SYBR Green).
- Capillary electrophoresis system (e.g., Fragment Analyzer).
Procedure:
- Transfection: Seed cells in a 24-well plate. At 70-80% confluency, co-transfect 200 ng of minigene plasmid with 50 nM of experimental or scrambled control ASO using Lipofectamine 3000 per manufacturer's protocol.
- RNA Isolation: 48 hours post-transfection, lyse cells in TRIzol. Isolate total RNA, treat with DNase I, and quantify.
- RT-PCR: Perform reverse transcription using random hexamers. Conduct PCR using primers spanning the minigene's constitutive exons.
- Product Analysis: Resolve PCR products via capillary electrophoresis. Quantify the percentage of transcripts including the target exon versus those skipping it using peak area integration.
Protocol 2: CRISPR/Cas9-Mediated Exon Inclusion for DMD Modeling/Correction
Objective: To permanently restore the reading frame in DMD patient-derived cells by excising a mutation-harboring exon via paired CRISPR/Cas9 cleavage. Materials:
- DMD patient-derived iPSCs or myoblasts with a known exon deletion (e.g., ΔExon 50).
- pX459 plasmids expressing SpCas9 and two sgRNAs targeting intronic regions flanking exon 51.
- Puromycin for selection.
- Nuclease-free water and PCR reagents.
- T7 Endonuclease I or next-generation sequencing (NGS) for indel analysis.
- Myogenic differentiation media.
Procedure:
- sgRNA Design & Cloning: Design two sgRNAs targeting conserved sequences in the introns immediately upstream and downstream of exon 51. Clone into pX459.
- Cell Transfection & Selection: Co-transfect both pX459 constructs into DMD cells using nucleofection. Apply puromycin (1-2 μg/mL) 48 hours post-transfection for 72 hours.
- Genomic DNA Analysis: Isolate genomic DNA from pooled edits or single-cell clones. Perform PCR across the target locus. Assess editing efficiency via T7E1 assay or NGS.
- RNA/Protein Validation: Differentiate edited iPSCs into myotubes. Isolate RNA and perform RT-PCR across the DMD target region. Analyze PCR products for exon 51 inclusion and correct frame. Confirm dystrophin expression by Western blot.
Protocol 3: Profiling Splicing Alterations in Cancer Cells with RNA-Seq
Objective: To identify genome-wide splicing changes in cancer cell lines harboring spliceosome mutations (e.g., SF3B1^mut). Materials:
- Isogenic pair of cancer cell lines (SF3B1 mutant vs. wild-type).
- TRIzol LS reagent.
- Poly(A) selection or rRNA depletion kit for RNA-seq library prep.
- High-throughput sequencer (Illumina platform).
- Computational tools: STAR aligner, rMATS, SUPPA2.
Procedure:
- RNA Extraction & QC: Extract high-quality total RNA (RIN > 8.5) from triplicate samples of each cell line.
- Library Preparation & Sequencing: Construct stranded RNA-seq libraries using poly(A) selection. Sequence on an Illumina NovaSeq to a depth of ~40-50 million paired-end 150bp reads per sample.
- Splicing Analysis: Align reads to the human reference genome (GRCh38) using STAR. Quantify splicing events (SE, MXE, A5SS, A3SS, RI) using rMATS with FDR < 0.05 and |ΔPSI| > 0.1 (where PSI = Percent Spliced In).
- Pathway Enrichment: Perform Gene Ontology (GO) and KEGG pathway analysis on genes harboring significant splicing alterations to identify dysregulated biological processes.
Visualizations
Diagram Title: Mechanism of Splicing Defects and Therapeutic Correction
Diagram Title: CRISPR/Cas9 Workflow for DMD Exon Editing
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Splicing Mutation Research
| Reagent / Material | Function in Research | Example Product / Assay |
|---|---|---|
| Splice-Switching ASOs | Chemically modified oligonucleotides to block splicing regulatory elements (ISE/ISS) and promote correct exon inclusion. | Morpholino oligos (e.g., for SMN2 exon 7 inclusion, Eteplirsen for DMD exon 51). |
| CRISPR/Cas9 Systems | For permanent genomic editing to correct splice site mutations, remove cryptic exons, or modulate splicing regulatory regions. | SpCas9 with paired sgRNAs for exon excision; base editors for splice site correction. |
| Minigene Splicing Reporters | Plasmid constructs containing genomic regions of interest to rapidly assay splicing patterns and test therapeutics in vitro. | pSpliceExpress or pMG2 vectors with cloned exons and flanking introns. |
| High-Throughput RNA-Seq Kits | For comprehensive, quantitative profiling of splicing isoforms and discovery of aberrant events in disease models. | Illumina Stranded mRNA Prep; NuGEN Ovation SoLo RNA-Seq System. |
| Splicing Analysis Software | Computational tools to identify and quantify differential splicing events from RNA-seq data. | rMATS, SUPPA2, LeafCutter for PSI (Ψ) calculation. |
| Isogenic Cell Line Pairs | Disease-relevant cell lines (e.g., iPSCs) with and without a specific splicing mutation, critical for controlled experimental comparison. | Patient-derived iPSCs corrected via CRISPR to create isogenic wild-type control. |
| In Vivo Splicing Reporters | Transgenic models expressing fluorescent proteins under control of alternative splicing events to visualize correction in real time. | Smn2-Δ7 reporter mouse; dual-fluorescence (e.g., GFP/RFP) minigene reporters. |
Within the context of a broader thesis on alternative splicing mutation research, CRISPR/Cas9 has emerged as an indispensable tool. Its precision, efficiency, and versatility enable researchers to model disease-associated splicing mutations, dissect regulatory elements, and develop potential therapeutic strategies with unprecedented control. This document provides detailed application notes and protocols for leveraging CRISPR/Cas9 in splicing research.
Core Applications and Quantitative Data
CRISPR/Cas9 is applied to splicing research through several key approaches, each with distinct efficiencies and purposes.
Table 1: Efficacy Metrics for CRISPR/Cas9 Applications in Splicing Mutation Research
| Application | Typical Target | Average Efficiency Range | Primary Readout | Key Benefit for Splicing Research |
|---|---|---|---|---|
| Exon Skipping/Inclusion | Splice Acceptor/Donor Sites | 20-60% (Indel Rate) | RT-PCR, RNA-Seq | Rapidly models aberrant splicing seen in disease. |
| Regulatory Element Disruption | Exonic/Intronic Splicing Enhancers/Silencers (ESEs, ISEs, ESSs, ISSs) | 30-70% (Editing Efficiency) | Splicing Reporter Assay, qRT-PCR | Functional mapping of splicing regulatory code. |
| Precise Mutation Knock-in | Disease-associated SNP at Splice Site | 5-30% (HDR Rate) | Sequencing, Functional Assays | Recapitulates patient-specific genetic variants. |
| Saturation Mutagenesis | Whole splice region | Varies by library size | High-throughput sequencing (NGS) | Comprehensive identification of cis-regulatory elements. |
| Gene Tagging (e.g., MS2, PP7) | Endogenous gene locus | 10-25% (Tagging Efficiency) | Live-cell imaging, Single-molecule RNA tracking | Visualizes transcription and co-transcriptional splicing dynamics. |
Detailed Experimental Protocols
Protocol 1: Disrupting a Canonical Splice Site to Induce Exon Skipping
Objective: To model a splicing defect by generating indels at a conserved GT/AG splice site via non-homologous end joining (NHEJ).
Materials:
- Cells: HEK293T or relevant disease-model cell line.
- CRISPR Components: Alt-R S.p. HiFi Cas9 Nuclease V3 (IDT), sgRNA targeting the splice site.
- Delivery: Lipofectamine CRISPRMAX Transfection Reagent.
- Analysis: RNA extraction kit, cDNA synthesis kit, PCR reagents, gel electrophoresis.
Method:
- Design sgRNA: Design a sgRNA targeting within 5 bp of the exon-intron boundary of the splice site (e.g., the last bases of an exon or first bases of an intron). Use tools like CHOPCHOP or CRISPick.
- Transfection: Complex 30 pmol of Cas9 protein with 45 pmol of sgRNA to form ribonucleoprotein (RNP). Transfect into 2e5 cells per well of a 24-well plate using Lipofectamine CRISPRMAX per manufacturer's protocol.
- Incubation: Culture cells for 72 hours to allow for editing and transcript turnover.
- Genomic Validation: Harvest a portion of cells for genomic DNA extraction. PCR-amplify the target region and perform T7 Endonuclease I assay or Sanger sequencing with tracking of indels by decomposition (TIDE) analysis to confirm editing efficiency.
- Splicing Analysis: Extract total RNA, synthesize cDNA, and perform RT-PCR with primers flanking the exon of interest. Analyze products by agarose gel electrophoresis. Successful splice site disruption will yield a smaller PCR product corresponding to the exon-skipped isoform.
- Quantification: Quantify band intensities using ImageJ software. Calculate percentage of exon skipping as: (Intensity of skipped product / Total intensity of all products) * 100.
Protocol 2: Precise Knock-in of a Splicing Mutation via HDR
Objective: To introduce a patient-specific single nucleotide variant (SNV) at a splice site using a single-stranded oligodeoxynucleotide (ssODN) donor template.
Materials:
- Cells: Cell line with high HDR efficiency (e.g., hPSCs, RPE1).
- CRISPR Components: As in Protocol 1.
- Donor Template: 200 nt ssODN homology-directed repair (HDR) template (Ultramer, IDT) containing the desired SNV, centrally located.
- Enhancers: Alt-R HDR Enhancer V2 (optional).
- Analysis: Next-generation sequencing (NGS) amplicon sequencing.
Method:
- Design: Design sgRNA with cut site as close as possible to the target SNV. Design the ssODN donor with ~90 nt homology arms on each side of the SNV. Incorporate silent blocking mutations in the sgRNA PAM/protospacer within the donor to prevent re-cutting.
- Transfection: Co-transfect RNP complex (as in Protocol 1) with 1 µL of 100 µM ssODN donor per well (24-well plate). Add HDR Enhancer if used.
- Enrichment (Optional): If using a fluorescent reporter or antibiotic resistance cassette linked to the repair, begin selection 48-72 hours post-transfection.
- Cloning & Screening: Single-cell clone the transfected population by limiting dilution. Allow 2-3 weeks for colony formation.
- Genotyping: Screen expanded clonal lines by PCR and Sanger sequencing of the target locus to identify correctly edited homozygous clones.
- Phenotypic Validation: Perform RT-PCR and RNA-seq on positive clones to confirm the alteration in splicing patterns caused by the precise mutation.
Visualizing Workflows and Pathways
Precise Splicing Mutation Knock-in via CRISPR HDR Workflow
Splicing Regulation by Enhancer (ESE) and Silencer (ESS) Elements
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for CRISPR-based Splicing Research
| Reagent / Material | Vendor Examples | Function in Splicing Mutation Research |
|---|---|---|
| High-Fidelity Cas9 Nuclease | IDT (Alt-R S.p. HiFi), NEB (HiFi Cas9) | Reduces off-target editing, crucial for accurate modeling of specific mutations. |
| Chemically Modified sgRNA | Synthego, IDT (Alt-R crRNA/tracrRNA) | Enhances stability and RNP formation efficiency, leading to higher editing rates. |
| HDR Donor Templates (ssODN) | IDT (Ultramer DNA Oligos), Twist Bioscience | Enables precise knock-in of single or multiple nucleotide changes to recreate patient variants. |
| Electroporation/Nucleofection Kits | Lonza (Nucleofector), Bio-Rad (Gene Pulser) | Efficient delivery of CRISPR components into hard-to-transfect primary or stem cells. |
| Splicing Reporter Minigene Vectors | Addgene (e.g., pSpliceExpress) | Validates the impact of cis-regulatory elements in a controlled context. |
| RT-PCR/qPCR Reagents for Isoforms | Bio-Rad, Thermo Fisher (TaqMan assays) | Quantifies relative expression of different mRNA splicing isoforms. |
| NGS Library Prep Kits (for RNA-seq) | Illumina (TruSeq Stranded mRNA), PacBio (Iso-Seq) | Provides comprehensive, unbiased analysis of global splicing changes. |
| T7 Endonuclease I / Mismatch Detection Kits | NEB, IDT | Rapidly assesses genomic editing efficiency at the target locus. |
| Single-Cell Cloning Dilution Plates | Corning, Thermo Fisher | Facilitates the isolation of genetically homogeneous isogenic clones post-editing. |
| Splicing Factor Antibodies (e.g., SRSF1, hnRNPA1) | Santa Cruz Biotechnology, Abcam | Validates protein-level changes or immunoprecipitates splicing complexes (RIP). |
Precision Editing in Practice: CRISPR/Cas9 Strategies to Model and Correct Splicing Defects
Application Notes
In the context of a CRISPR/Cas9-based thesis aimed at creating or correcting disease-relevant alternative splicing mutations, precise targeting of core splice elements is paramount. The design principles focus on disrupting or restoring the canonical splicing code to alter exon inclusion rates. The efficacy of such interventions is quantified by metrics like Percent Spliced In (PSI) and is highly dependent on the targeted element's sequence context and predicted strength.
Key Quantitative Data on Splice Site Targeting Outcomes
Table 1: Representative CRISPR/Cas9-Mediated Splicing Modulation Outcomes
| Targeted Element | Exon Context | Average ΔPSI Reported | Key Determinant of Efficacy | Primary Assay |
|---|---|---|---|---|
| Splice Donor Site (GT) | Cassette Exon | -40% to +85%* | Proximity to AG-rich region; Strength of competitor site | RT-PCR, RNA-seq |
| Splice Acceptor Site (AG) | Cassette Exon | -35% to +75%* | Pyrimidine tract length; Branch point distance | RT-PCR, RNA-seq |
| Exonic Splicing Enhancer (ESE) | Alternative Exon | -15% to -60% | SR protein binding motif score; Redundancy of motifs | RT-PCR, Minigene |
| Intronic Splicing Silencer (ISS) | Alternative Exon | +20% to +50% | hnRNP protein binding affinity; Secondary structure | RT-PCR, RNA-seq |
| Intronic Splicing Enhancer (ISE) | Alternative Exon | -25% to -55% | Proximity to splice site; Density of regulatory motifs | Minigene, RNA-seq |
*Positive ΔPSI indicates exon inclusion increase, negative indicates decrease. Outcomes vary dramatically based on creation/restoration of a functional vs. non-functional site.
Experimental Protocols
Protocol 1: Design and Validation of sgRNAs Targeting Splice Elements
Objective: To induce exon skipping or inclusion by disrupting splice donor/acceptor sites or regulatory sequences (ESEs, ISEs).
Materials:
- sgRNA Design Tools: CRISPick, ChopChop, SpliceRush.
- Prediction Algorithms: ESEfinder (v3.0), RESCUE-ESE, FAS-ESS, SpliceAid2.
- Target Genomic DNA
- Cloning reagents: pSpCas9(BB)-2A-Puro (PX459) V2.0 vector, BbsI restriction enzyme, T4 DNA ligase.
- Cell Line: HEK293T or relevant disease model cell line.
- Transfection reagent: Lipofectamine 3000.
- Validation Primers flanking the alternative exon.
Procedure:
- Target Identification: Using human genome assembly GRCh38, identify the target exon and its flanking introns.
- Element Mapping: Use ESEfinder and SpliceAid2 to map putative ESE (e.g., SF2/ASF, SC35 motifs) and ISE motifs within the target exon and adjacent introns (approx. 300bp).
- sgRNA Design:
- For donor/acceptor disruption: Design two sgRNAs bracketing the canonical GT/AG dinucleotide. Place the Cas9 cut site 1-3 bp upstream of the dinucleotide to promote error-prone NHEJ that disrupts it.
- For ESE/ISE disruption: Design sgRNAs with protospacers directly overlapping the highest-scoring predicted SR protein binding motif(s). Prioritize motifs with low redundancy.
- Cloning into PX459: Digest vector with BbsI, anneal oligos encoding the sgRNA, and ligate following the Zhang Lab protocol.
- Cell Transfection: Seed cells at 70% confluence in 6-well plates. Co-transfect 1.5 µg of each sgRNA plasmid using Lipofectamine 3000.
- Harvest and Analysis: 48-72h post-transfection, harvest total RNA with TRIzol, perform DNase I treatment, and synthesize cDNA.
- Splicing Analysis: Perform RT-PCR with primers in the constitutive exons. Analyze products via capillary electrophoresis (e.g., QIAxcel) or agarose gel electrophoresis to quantify exon inclusion/exclusion. Calculate PSI = (Inclusion peak area / (Inclusion + Exclusion peak area)) * 100.
Protocol 2: Minigene Splicing Reporter Assay for Validating Element Function
Objective: To functionally validate the role of a predicted ESE or ISE element before CRISPR editing in the endogenous locus.
Materials:
- Cloning Vector: pSpliceExpress or analogous exon-trapping vector.
- PCR Reagents with high-fidelity polymerase.
- Restriction Enzymes (e.g., BamHI, XhoI).
- Cell Line: HEK293T (for robust splicing).
- Dual-Luciferase Reporter Assay System (optional, for quantification).
Procedure:
- Amplify Genomic Region: PCR amplify a genomic fragment containing the target alternative exon (~300bp upstream to ~300bp downstream) from wild-type DNA.
- Clone into Reporter: Insert the fragment into the multicoloring site of the minigene vector between two constitutive exons.
- Site-Directed Mutagenesis: Generate mutant constructs where the core ESE/ISE motif (e.g., GGAGG → GGAtt) is disrupted using overlap extension PCR.
- Transfection: Transfect wild-type and mutant minigene constructs into HEK293T cells in triplicate.
- RNA Isolation & RT-PCR: 24h post-transfection, isolate RNA and perform RT-PCR using vector-specific primers flanking the inserted region.
- Quantification: Analyze PCR products by gel electrophoresis. Compare the ratio of spliced isoforms (exon included vs. skipped) between wild-type and mutant constructs. A significant change confirms the element's functional role.
Visualizations
Title: CRISPR Splicing Modulation Experimental Workflow
Title: Splicing Code Elements Determine Exon Inclusion vs Skipping
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Splice-Targeting CRISPR Experiments
| Reagent / Material | Function & Rationale | Example Product / Vendor |
|---|---|---|
| High-Specificity Cas9 Nuclease | Enables precise DSB induction with minimal off-target effects, critical near highly conserved splice sites. | Alt-R S.p. HiFi Cas9 Nuclease V3 (IDT) |
| Chemically Modified sgRNAs | Enhances stability and cutting efficiency; crucial for targeting nucleosome-dense regions like intron-exon junctions. | Synthego sgRNA EZ Kit |
| Splice-Sensitive RT-PCR Kit | Allows accurate quantification of alternative splicing isoforms from low-input RNA samples post-editing. | TaqMan Alternative Splicing Assays (Thermo Fisher) |
| Next-Gen Sequencing Library Prep for Splicing | Captures genome-wide on-target editing and off-target splicing effects. | KAPA RNA HyperPrep Kit with RiboErase (Roche) |
| Minigene Splicing Reporter Vector | For rapid functional validation of putative ESE/ISE elements prior to endogenous editing. | pSpliceExpress (Addgene #52857) |
| Splice-Junction-Focused Analysis Software | Quantifies PSI from RNA-seq data and identifies novel or cryptic splicing events. | rMATS, MAJIQ, SpliceBERT |
This document provides Application Notes and Protocols for using CRISPR/Cas9 variants in the study and correction of disease-causing splicing mutations. Within the broader thesis on CRISPR/Cas9 alternative splicing mutation research, these tools enable precise dissection and repair of cis-acting regulatory elements—such as splice donor/acceptor sites, branch points, and exonic/intronic splicing enhancers/silencers (ESEs/ISEs, ESSs/ISSs)—offering novel therapeutic avenues for genetic disorders.
Quantitative Comparison of Cas9 Variants for Splicing Modulation
Table 1: Efficacy and Outcomes of Cas9 Variants in Splicing Correction
| Cas9 Variant | Primary Action | Typical Target (for Splicing) | Editing Outcome | Average Correction Efficiency (Reported Range) | Primary Indels/By-products |
|---|---|---|---|---|---|
| Wild-Type Nuclease | Creates DSBs | Exon-Intron Junctions, Splicing Regulators | Exon skipping/inclusion via NHEJ-mediated indels | 15-50% (NHEJ-dependent) | High (>30% frameshift indels) |
| D10A Nickase (nCas9) | Creates single-strand nicks | Pairs targeting cis-elements | Preciser small deletions/insertions via HDR or MMEJ | 5-25% (HDR-dependent) | Reduced; paired nicks can create DSBs |
| BE3 (C→T, G→A) Base Editor | C•G to T•A conversion | Point mutations in splice sites or ESEs/ESSs | Direct point mutation to restore or disrupt motifs | 30-70% (in transfected cells) | Very low (<1% indels); bystander edits possible |
| ABE8e (A→G, T→C) Base Editor | A•T to G•C conversion | Point mutations in splice sites or ESEs/ESSs | Direct point mutation to restore or disrupt motifs | 40-80% (in transfected cells) | Very low (<1% indels); bystander edits possible |
Table 2: Key Considerations for Tool Selection in Splicing Research
| Factor | Nuclease | Nickase | Base Editor |
|---|---|---|---|
| Goal | Complete disruption/removal of regulatory element | Subtler disruption or small sequence alteration | Single-nucleotide conversion to correct/alter motif |
| Risk of Genomic Toxicity | High (off-target DSBs) | Moderate | Low (no DSBs) |
| HDR Requirement | Yes for precise correction | Yes for precise insertion | No |
| Applicable Mutations | Large deletions, exon skipping | Small deletions/insertions | Point mutations at splice sites, ESEs, ESSs |
Experimental Protocols
Protocol 1: Disrupting an Aberrant Splice Acceptor Site Using Cas9 Nuclease
Objective: Induce frameshift indels via NHEJ to permanently inactivate a mutated splice acceptor site, forcing skipping of a mutant exon. Materials: See "Scientist's Toolkit" below. Procedure:
- sgRNA Design: Design two sgRNAs flanking the aberrant splice acceptor site within the intron (e.g., -5 to -30 bp upstream of the exon).
- RNP Complex Formation: Complex 10 µg of purified SpCas9 protein with 5 µg of each sgRNA (total 10 µg sgRNA) in PBS+ buffer. Incubate at 25°C for 10 min.
- Cell Nucleofection: Harvest 1x10^6 patient-derived fibroblasts (e.g., from a SMA or DMD model). Resuspend cell pellet in 100 µL Nucleofector Solution. Mix with RNP complex. Electroporate using a 4D-Nucleofector (program: CM-137). Immediately add pre-warmed medium.
- Analysis (72 hrs post):
- Genomic DNA: Extract gDNA. PCR amplify target region. Analyze indel frequency via T7 Endonuclease I assay or next-generation sequencing (NGS).
- RNA: Extract total RNA. Perform RT-PCR across the targeted exon. Analyze products via agarose gel electrophoresis for exon skipping.
- Validation: Sanger sequence cDNA clones to confirm novel splicing patterns.
Protocol 2: Correcting a Point Mutation in an Exonic Splicing Enhancer Using ABE8e
Objective: Convert a disease-causing A•T to G•C point mutation within an ESE to restore correct splicing. Materials: See "Scientist's Toolkit" below. Procedure:
- sgRNA Design: Design an sgRNA positioning the target A (on the non-template strand) within the editing window (positions 4-8, counting PAM as 21-23) of ABE8e.
- Plasmid Transfection: Co-transfect HEK293T cells (or relevant disease-model iPSCs) in a 24-well plate with 500 ng of ABE8e expression plasmid (e.g., pCMV_ABE8e) and 250 ng of sgRNA expression plasmid (e.g., pU6-sgRNA) using 1.5 µL of lipofectamine 3000.
- Cell Harvest (Day 5): Harvest cells for gDNA and RNA extraction.
- Analysis:
- Editing Efficiency: Amplify target region from gDNA. Perform deep sequencing (NGS) to quantify A-to-G conversion efficiency and bystander edits.
- Splicing Assay: Perform RT-PCR from extracted RNA. Resolve products on a high-percentage agarose gel (3%) or via capillary electrophoresis (e.g., Bioanalyzer) to quantify restoration of correct splicing isoforms.
- Functional Assay: Perform a disease-relevant functional assay (e.g., ELISA for protein, contractility for cardiomyocytes).
Visualizations
Diagram 1: CRISPR Toolbox Action on Splicing Regulatory Elements
Diagram 2: Base Editor Correction of an Exonic Splicing Enhancer (ESE) Mutation Workflow
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Materials for Splicing-Focused CRISPR Experiments
| Reagent/Material | Supplier Examples | Function in Splicing Research |
|---|---|---|
| SpCas9 Nuclease (WT) | Thermo Fisher, Synthego | Creates DSBs for disruptive editing of splice sites. |
| HiFi Cas9 Protein | IDT, Thermo Fisher | High-fidelity variant for reduced off-target DSB generation. |
| D10A (nCas9) Expression Plasmid | Addgene (#41816) | Enables paired nicking for precise small edits in regulatory elements. |
| ABE8e & BE4max Plasmids | Addgene (#138489, #139999) | For efficient single-base conversion to correct point mutations in ESEs/ISEs or splice sites. |
| Chemically Modified sgRNAs (synthego) | Synthego, IDT | Enhanced stability and editing efficiency for RNP delivery. |
| 4D-Nucleofector System & Kits | Lonza | High-efficiency delivery of RNPs into primary cells and iPSCs. |
| Lipofectamine 3000 | Thermo Fisher | Standard transfection reagent for plasmid delivery in cell lines. |
| T7 Endonuclease I | NEB | Rapid detection of indel formation after nuclease treatment. |
| Agilent 2100 Bioanalyzer & RNA Kits | Agilent | High-resolution analysis of splicing isoform changes post-editing. |
| SMARTer PCR cDNA Synthesis Kit | Takara Bio | Efficient cDNA synthesis for RT-PCR analysis of low-abundance isoforms. |
Within the broader thesis investigating CRISPR/Cas9 applications for correcting disease-causing alternative splicing mutations, this application note details practical methodologies for targeted exon skipping and inclusion. These approaches aim to reframe mutated or in-frame exons, offering therapeutic strategies for genetic disorders like Duchenne Muscular Dystrophy (DMD) and Spinal Muscular Atrophy (SMA).
Conventional CRISPR/Cas9-mediated gene knockout relies on frameshift indels from non-homologous end joining (NHEJ). For splicing modulation, the strategy is refined: paired single-guide RNAs (sgRNAs) are designed to target flanking intronic regions of a specific exon. Dual double-strand breaks (DSBs) induce deletion of the intervening exon via microhomology-mediated end joining (MMEJ) or alt-EJ, or its inversion via NHEJ-mediated rearrangement, effectively skipping it from the mature mRNA. For exon inclusion, CRISPRa/i systems can modulate splicing enhancer/silencer elements.
Table 1: Efficacy of CRISPR-Mediated Exon Skipping in Pre-Clinical Models
| Disease Model (Exon Target) | Delivery Method | Skipping Efficiency (RT-PCR) | Functional Protein Rescue | Citation (Year) |
|---|---|---|---|---|
| DMD (mdx mouse, Exon 23) | AAV9-Cas9/sgRNA | 2-60% (muscle) | Dystrophin+: Up to 80% fibers | Nelson et al., 2019 |
| DMD (human cardiomyocytes, Exon 51) | RNP electroporation | ~90% (in vitro) | Dystrophin detected | Moretti et al., 2022 |
| SMA (IPS cells, SMN2 Exon 7) | Lentiviral SaCas9 | ~60% inclusion | Increased SMN protein | Kim et al., 2023 |
| CFTR (F508del, Exon 11)* | Adenoviral delivery | ~10-20% correction | Partial CFTR function | Recent studies |
*CFTR example often involves ablated exon removal to restore reading frame.
Table 2: Comparison of CRISPR Systems for Splicing Modulation
| System | Nuclease Activity | Primary Use for Splicing | Key Advantage | Key Limitation |
|---|---|---|---|---|
| SpCas9 (NHEJ) | DSB | Exon deletion/inversion via dual-cut | High efficiency, well-characterized | Off-target DSB risk, large size |
| Cas9 nickase | Single-strand nick | Paired nicks for exon deletion | Reduced off-target indels | Lower deletion efficiency |
| dCas9 (CRISPRi) | Catalytically dead | Block splicing silencer elements | No DSBs, reversible | Requires sustained expression |
| dCas9-VPR (CRISPRa) | Catalytically dead | Activate splicing enhancers | No DSBs, can promote inclusion | Potential transcriptional noise |
Experimental Protocols
Protocol 1: Dual-sgRNA Design & Validation for Exon Deletion
Objective: To design and validate sgRNA pairs that efficiently delete a target exon.
- Design:
- Identify target exon coordinates (hg38/mm10). Select unique 20-nt sgRNA sequences within flanking introns (50-500 bp from exon-intron junctions).
- Use tools (CHOPCHOP, CRISPick) to predict on-target efficiency and minimize off-targets.
- Order: Chemically synthesize two sgRNAs with tracrRNA scaffold or as single-guide RNAs.
- In Vitro Validation (HEK293T cells):
- Transfection: Co-transfect 500 ng SpCas9 expression plasmid + 125 ng of each sgRNA plasmid (or 50 pmol total sgRNA as RNP) per well in a 24-well plate.
- Genomic DNA Extraction: 72 hrs post-transfection, extract gDNA. Perform PCR across the targeted region (amplicon spanning both cut sites).
- Analysis: Run PCR product on agarose gel. Successful deletion yields a smaller band. Confirm by Sanger sequencing. Quantify deletion efficiency using TIDE or ICE analysis.
Protocol 2: AAV-Mediated Delivery forIn VivoExon Skipping (Mouse Model)
Objective: To achieve body-wide or tissue-specific exon skipping in a murine disease model.
- Vector Production: Package a single AAV vector expressing SaCas9 (smaller than SpCas9) and a dual-sgRNA expression cassette under a tissue-specific promoter (e.g., MHCK7 for muscle) using the AAV9 serotype.
- Animal Injection: Inject neonatal (P1) or adult mice via intravenous (retro-orbital) or intramuscular route. Standard dose: 1x10^14 – 5x10^14 vg/kg for systemic delivery.
- Tissue Analysis (4-8 weeks post-injection):
- Genomic: Isolate DNA from harvested tissues. Perform PCR/sequencing to detect exon deletion.
- Transcriptomic: Isolate RNA, perform RT-PCR across the spliced region. Quantify exon-skipped mRNA % using capillary electrophoresis (Fragment Analyzer).
- Proteomic: Perform western blot or immunohistochemistry for the rescued protein.
Protocol 3: dCas9-Based Splicing Modulator Screening (CRISPRi/a)
Objective: To identify functional splicing regulatory elements (SREs) for exon inclusion.
- Library Design: Design sgRNA tiling library targeting ~200 bp upstream/downstream of target exon, focusing on putative SRE motifs (e.g., for silencing: hnRNP A1, for enhancing: SR protein sites).
- Screening: Lentivirally transduce cells stably expressing dCas9-KRAB (for i) or dCas9-VPR (for a) with the sgRNA library at low MOI.
- Selection & Sequencing: After 7-14 days, isolate genomic DNA, PCR amplify sgRNA region, and sequence via NGS. Enriched/depleted sgRNAs indicate regulatory SREs affecting exon inclusion (measured via concomitant RNA-seq).
Visualization
CRISPR Workflow for Exon Skipping
Mechanism of Exon Deletion Therapy
The Scientist's Toolkit
Table 3: Essential Research Reagents & Solutions
| Item | Function & Role in Experiment | Example/Note |
|---|---|---|
| High-Fidelity Cas9 Nuclease | Catalyzes DSB at genomic target. Purified protein for RNP formation reduces off-targets and immune response. | SpCas9, SaCas9 (for AAV packaging). |
| Chemically Modified sgRNA | Guides Cas9 to target sequence. Chemical modifications (2'-O-methyl, phosphorothioate) enhance stability in vivo. | Synthesized via solid-phase, HPLC-purified. |
| AAV Vector (Serotype 9) | In vivo delivery vehicle. AAV9 shows broad tissue tropism (muscle, heart, CNS). Single or dual-vector systems. | Must keep cargo < ~4.7 kb. |
| Nucleofection/Electroporation Kit | For efficient RNP or plasmid delivery into hard-to-transfect primary cells (e.g., myoblasts, iPSCs). | Lonza Nucleofector, Neon System. |
| T7 Endonuclease I or Surveyor Assay | Detects Cas9-induced indels via mismatch cleavage of heteroduplex DNA. Quick validation of editing efficiency. | Less sensitive than NGS but rapid. |
| Next-Gen Sequencing (NGS) Library Prep Kit | For unbiased quantification of editing outcomes (deletion %), splice variants, and off-target profiling. | Illumina MiSeq, amplicon-seq. |
| Splice-Sensitive RT-PCR Reagents | Detects changes in mRNA splicing patterns. Use primers in exons flanking the target. | Analysis via agarose gel or Bioanalyzer. |
| Antibody for Target Protein | Validates functional rescue at protein level via western blot or IHC. Critical for preclinical studies. | e.g., Dystrophin (MANDYS106), SMN. |
Within the broader thesis exploring CRISPR/Cas9 applications for correcting disease-causing alternative splicing mutations, the selection and optimization of a delivery vehicle is a critical determinant of therapeutic success. This document provides Application Notes and detailed Protocols for three leading platforms: Adeno-Associated Virus (AAV), Lipid Nanoparticles (LNPs), and Ribonucleoprotein (RNP) complexes. Each system presents unique trade-offs between payload capacity, immunogenicity, durability, and manufacturability, which must be evaluated for splice-targeting gene editors.
Application Notes: Platform Comparison
Table 1: Quantitative Comparison of Delivery Platforms for Splice-Correction
| Parameter | AAV | Lipid Nanoparticles (LNPs) | RNP Complexes |
|---|---|---|---|
| Typical Payload | DNA (ssAAV ~4.7 kb, scAAV ~2.3 kb) | mRNA/sgRNA or DNA (Plasmid) | Pre-assembled Cas9 protein + sgRNA |
| Delivery Efficiency (In Vitro, %) | High in permissive cells (60-95%) | Very High in many cell types (70-95%) | High (80-95%) |
| In Vivo Tropism | Defined by serotype; can be engineered | Broad; can be targeted with ligands | Local delivery (e.g., electroporation) |
| Onset of Action | Slow (weeks for peak expression) | Fast (hours to days) | Fastest (immediate, hours) |
| Duration of Effect | Long-term (potentially years) | Transient (days to weeks) | Ultra-short (days) |
| Immunogenicity Risk | High (pre-existing/vector immunity) | Moderate (lipid/reactogenicity) | Lowest (no DNA, short exposure) |
| Payload Capacity | Limited | High/Large | Limited to protein complex size |
| Manufacturing Scalability | Complex, expensive | Easier, scalable | Complex protein production |
| Key Risk for Splicing | Genomic integration risk (rare), long-term off-target exposure | Cytotoxicity at high doses, transient expression may require redosing | Rapid degradation may limit efficacy in post-mitotic tissues |
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function & Relevance |
|---|---|
| AAV Serotype Library (e.g., AAV1, AAV2, AAV5, AAV6, AAV8, AAV9, AAV-DJ, AAV-PHP.eB) | Enables tropism screening for target tissue (e.g., CNS, muscle, liver) in vitro and in vivo. |
| Ionizable Cationic Lipid (e.g., DLin-MC3-DMA, SM-102, ALC-0315) | Critical LNP component for encapsulating nucleic acids and enabling endosomal escape. |
| Purified Cas9 Nuclease (WT or HiFi) | Essential for forming RNP complexes; high-fidelity variants reduce off-target editing in splicing correction. |
| Chemically Modified sgRNA | Enhances stability and reduces immunogenicity for LNP and RNP delivery; critical for in vivo efficacy. |
| Splicing Reporter Cell Line (e.g., with mutant GFP/luciferase minigene) | Functional assay to quantify splice-correction efficiency (e.g., restoration of fluorescence/luminescence). |
| Next-Generation Sequencing (NGS) Kit for RNA-seq or Targeted Amplicon-seq | Gold standard for assessing splicing outcome changes (exon inclusion/exclusion) and transcriptome-wide off-target effects. |
Experimental Protocols
Protocol 1: AAV-Mediated Delivery of SaCas9/sgRNA for Exon Inclusion Objective: To package a splice-switching SaCas9 (with smaller size than SpCas9) and sgRNA expression cassette into AAV for durable correction of exon-skipping mutations.
Materials: pAAV-saCas9-U6-sgRNA (therapeutic construct), pHelper, pAAV-RC (serotype of choice), HEK293T cells, PEI-Max, Opti-MEM, PBS-MK (PBS with 1mM MgCl2, 2.5mM KCl), Benzonase, Iodixanol gradient solutions, Amicon Ultra-15 centrifugal filters.
Method:
- Triple Transfection: Seed HEK293T cells in 15-cm dishes. At 70-80% confluency, co-transfect using PEI-Max (1:3 DNA:PEI ratio): 7 µg pAAV-saCas9-U6-sgRNA, 14 µg pHelper, and 7 µg pAAV-RC in Opti-MEM.
- Harvest: 72 hours post-transfection, collect cells and media. Freeze-thaw lysate 3x, treat with 50 U/mL Benzonase (37°C, 30 min) to degrade unpackaged DNA.
- Purification: Clarify lysate by centrifugation. Layer supernatant on a pre-formed iodixanol step gradient (15%, 25%, 40%, 60%) in a sealed tube. Ultracentrifuge at 350,000 x g for 1.5 hours.
- Collection & Concentration: Extract the opaque 40% iodixanol fraction containing purified AAV. Concentrate and buffer exchange into PBS-MK using a 100kDa Amicon filter. Titrate via qPCR.
- In Vitro Validation: Transduce target cells (e.g., patient-derived fibroblasts) at an MOI of 10^4-10^5 vg/cell. After 7-14 days, harvest RNA for RT-PCR analysis of corrected splicing.
Protocol 2: LNP Formulation of Cas9 mRNA and sgRNA for Hepatocyte Splice-Correction Objective: To formulate ionizable LNPs encapsulating Cas9 mRNA and chemically modified sgRNA for in vivo delivery to the liver.
Materials: Cas9 mRNA (modified, e.g., 5-mC, Ψ), sgRNA (chemically modified with 2'-O-methyl, phosphorothioate), Ionizable lipid (e.g., SM-102), DSPC, Cholesterol, PEG-lipid, Acetic acid buffer (pH 4.0), Ethanol, TFF system, PBS.
Method:
- Lipid Preparation: Dissolve ionizable lipid, DSPC, cholesterol, and PEG-lipid (molar ratio 50:10:38.5:1.5) in ethanol.
- Aqueous Phase Preparation: Dilute Cas9 mRNA and sgRNA (mass ratio ~5:1) in acidic acetate buffer (pH 4.0).
- Microfluidic Mixing: Use a microfluidic device (e.g., NanoAssemblr) to rapidly mix the aqueous and ethanol phases at a 3:1 flow rate ratio (aqueous:ethanol). Total flow rate ~12 mL/min.
- Buffer Exchange & Purification: Dilute the formed LNP suspension 1:5 in PBS. Concentrate and dialyze against PBS using Tangential Flow Filtration (TFF) with a 100kDa membrane to remove ethanol and residual buffer.
- Characterization & Delivery: Measure particle size (target 70-100 nm) via DLS and encapsulation efficiency (>90%) by RiboGreen assay. Administer intravenously to mouse models at 1-3 mg mRNA/kg dose. Assess liver splicing correction via RNA-seq 3-7 days post-injection.
Protocol 3: Electroporation of RNP Complexes for Excision of a Pseudoexon Objective: To deliver pre-complexed Cas9 protein and sgRNA via nucleofection for rapid, DNA-free correction of a deep-intronic mutation creating a cryptic splice site.
Materials: Recombinant Cas9 protein (Alt-R S.p. HiFi Cas9), Alt-R CRISPR-Cas9 crRNA (target-specific) & tracrRNA (modified), Nuclease-Free Duplex Buffer, Nucleofector Device & appropriate Kit (e.g., P3 for primary cells), pre-warmed culture medium.
Method:
- RNP Complex Assembly: Resuspend crRNA and tracrRNA to 100 µM in duplex buffer. Mix equal volumes, heat at 95°C for 5 min, cool to RT to form 50 µM sgRNA. Incubate 60 pmol Cas9 protein with 72 pmol sgRNA (1.2:1 molar ratio) in a total volume of ~10 µL for 10-20 min at room temperature.
- Cell Preparation: Harvest 0.5-1.0 x 10^5 target cells (e.g., CD34+ HSPCs or myoblasts) per reaction. Wash with PBS and resuspend in Nucleofector Solution.
- Nucleofection: Mix cell suspension with pre-assembled RNP complex. Transfer to a certified cuvette. Run the appropriate Nucleofector program (e.g., EO-100 for HSPCs).
- Recovery & Analysis: Immediately add pre-warmed medium to cuvette. Transfer cells to a culture plate. Analyze editing and splicing outcomes 48-72 hours post-electroporation via targeted amplicon sequencing (for indels) and RT-PCR (for splicing).
Visualizations
Title: Therapeutic Development Workflow for Splice-Correction
Title: Intracellular Delivery Pathways of AAV, LNP, and RNP
Overcoming Challenges: Optimizing CRISPR Efficiency and Specificity for Splicing Loci
Application Notes Within CRISPR/Cas9 research aimed at modeling alternative splicing mutations via targeted intronic or exonic disruptions, repetitive genomic regions present a significant challenge. These areas, including satellite DNA, transposable elements, and multi-copy gene families, are hotspots for off-target editing due to homologous sequences that can be recognized by guide RNAs (gRNAs) with imperfect complementarity. Off-target effects in these regions can lead to chromosomal rearrangements, aneuploidy, and transcriptional dysregulation, confounding phenotypic analyses in splicing mutation studies. Recent data (2023-2024) underscores the severity of this issue:
Table 1: Quantified Risk of Off-Target Effects in Repetitive Regions
| Metric | Low-Complexity Region (e.g., Alu) | Telomeric/Subtelomeric Repeat | Ribosomal DNA (rDNA) Array |
|---|---|---|---|
| Typical # of Genomic Loci | ~1.1 million (Alu) | 46-92 telomeres | ~300-400 copies |
| Reported Off-Target Rate* | Up to 15% higher than unique sites | 8-12% observed indel frequency | >20% variability in copy number post-editing |
| Primary Consequence | Non-specific indels, genomic instability | Telomere length alteration, end-to-end fusions | rDNA copy number change, nucleolar stress |
| Key Detection Method | WGS or targeted long-read sequencing | Telomere-specific FISH, qPCR | ddPCR, Southern blot |
*Compared to perfectly matched on-target site in a unique genomic region.
Experimental Protocol: Assessing Off-Target Effects in a Repetitive Region for a Splicing Mutation Model
Objective: To evaluate the specificity of a Cas9/gRNA complex designed to disrupt a splice acceptor site within a gene of interest that has paralogous sequences in the genome.
Materials:
- Cell Line: Target human iPSCs or cell line.
- CRISPR Components: SpCas9 nuclease, on-target gRNA (designed against the intended splice site).
- Control: Non-targeting gRNA.
- Transfection Reagent: (e.g., Lipofectamine CRISPRMAX).
- Analysis Reagents: Lysis buffer, PCR primers for on-target and predicted off-target loci, NGS library prep kit, T7 Endonuclease I (T7EI).
Procedure:
- In Silico Prediction: Use Cas-OFFinder and UCSC Genome Browser to identify all genomic loci with ≤5 mismatches to the on-target gRNA, with emphasis on repetitive families.
- Cell Transfection: Transfect cells with SpCas9 + on-target gRNA complex. Include a non-transfected control.
- Genomic DNA Harvest: 72 hours post-transfection, harvest and extract gDNA.
- Primary On-Target Efficiency Check:
- Amplify the on-target genomic region (500-600 bp amplicon).
- Perform T7EI assay on the purified PCR product.
- Run digested products on an agarose gel; quantify cleavage efficiency.
- Comprehensive Off-Target Analysis:
- For predicted sites (≤5 mismatches): Perform PCR amplification of each potential off-target locus from the harvested gDNA.
- For genome-wide unbiased discovery: Prepare an NGS library from the transfected pool gDNA using a kit like GUIDE-seq or CIRCLE-seq, following the manufacturer's protocol. Sequence on an Illumina platform.
- Analyze NGS data with pipelines (e.g., CRISPResso2) to quantify indel frequencies at all aligned loci.
- Validation: Top 5-10 candidate off-target sites (including those in repetitive zones) from NGS data must be validated by targeted amplicon sequencing from independently transfected cell pools.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Off-Target Analysis in Repetitive Regions
| Item | Function | Example Product/Assay |
|---|---|---|
| High-Fidelity Cas9 Variant | Reduces tolerance to gRNA mismatches, lowering off-target activity. | SpCas9-HF1, eSpCas9(1.1) |
| Chemically Modified gRNA | Improves stability and can enhance specificity by reducing affinity for off-target sequences. | 2'-O-methyl 3' phosphorothioate modified crRNA/tracrRNA |
| Unbiased Off-Target Discovery Kit | Identifies genome-wide double-strand breaks without prior sequence bias, critical for repeats. | GUIDE-seq, CIRCLE-seq |
| Long-Read Sequencer | Accurately maps edits within highly identical repetitive sequences. | PacBio Sequel II, Oxford Nanopore MinION |
| Digital PCR System | Absolutely quantifies copy number variations in repetitive arrays post-editing. | ddPCR for rDNA copy number |
| Bioinformatics Pipeline | Aligns sequencing reads to repetitive regions and calls variants with high confidence. | CRISPResso2, WGS + RepeatMasker |
Visualizations
Diagram 1: Off-target analysis workflow for splicing studies.
Diagram 2: Impact of repeat off-targets on splicing research.
(Framed within a thesis on CRISPR/Cas9 alternative splicing mutation research)
1. Introduction Within the broader objective of generating precise alternative splicing mutations to model disease or correct pathogenic isoforms, the design of single-guide RNAs (gRNAs) targeting splicing regulatory sequences (SRS) presents unique challenges. Unlike coding exons, SRS—including splice donors, acceptors, branch points, and exonic/intronic splicing enhancers/silencers (ESEs, ISEs, ESSs, ISSs)—are degenerate and context-dependent. This document provides application notes and optimized protocols for designing and validating gRNAs to disrupt or create these regulatory elements using CRISPR/Cas9.
2. Quantitative Parameters for SRS-Targeting gRNA Design Optimal gRNA design balances general CRISPR efficiency rules with SRS-specific localization. Key quantitative parameters, derived from recent literature and tool benchmarks, are summarized below.
Table 1: Prioritization Metrics for SRS-Targeting gRNA Design
| Parameter | Optimal Target/Score | Rationale & Tool for Calculation |
|---|---|---|
| On-Target Efficiency Score | >60 (Doench2016 score) | Predicts Cas9 cutting likelihood. Use Azimuth or CRISPick. |
| Off-Target Potential | Zero perfect matches in genome; <3 mismatches in exonic/noncoding conserved regions. | Minimizes unintended genomic edits. Use CRISPOR or Cas-OFFinder. |
| Distance to SRS Core | ≤10 bp from essential regulatory nucleotide (e.g., GT/AG dinucleotide). | Maximizes probability of disrupting SRS function via indel. |
| SRS Functional Score | High-confidence annotation from >2 databases (e.g., SpliceAid2, ESEfinder, HSF). |
Ensures target is a validated regulatory element. |
| gRNA Positioning | Prefer strand opposite to splicing factor binding site. | May enhance disruption by targeting the binding motif directly. |
| Local GC Content | 40-60% | Affects gRNA stability and Cas9 activity. |
Table 2: Comparison of SRS Annotation Resources
| Database/Tool | SRS Types Annotated | Organism | Key Feature |
|---|---|---|---|
SpliceAid 3 |
ESE, ESS, ISE, ISS, binding proteins | Human, Mouse | Experimental and predicted data. |
Human Splicing Finder (HSF 3.1) |
Donor, Acceptor, Branch Point, ESE/ESS | Human | Integrates multiple algorithms. |
SpliceDB |
Splice sites | 14 species | Curated from RNA-seq data. |
DeepSEA |
Epigenomic & splicing effect predictions | Human | Computational prediction of variant effects. |
3. Detailed Experimental Protocol: gRNA Validation for Splicing Disruption
A. Protocol: In Vitro Splicing Reporter Assay Objective: Functionally validate gRNA efficacy in disrupting an SRS prior to genomic editing. Materials: See "Scientist's Toolkit" below. Workflow: 1. Cloning: Subclone the genomic region of interest (containing the wild-type SRS) into a mammalian splicing reporter vector (e.g., pSpliceExpress). 2. gRNA Constructs: Clone top 2-3 candidate gRNA sequences into a Cas9/sgRNA expression plasmid (e.g., pX330). 3. Co-transfection: In a 24-well plate, co-transfect HEK293T cells with: * 400 ng splicing reporter plasmid. * 200 ng pX330-gRNA plasmid. * 100 ng fluorescent transfection control plasmid (e.g., pMaxGFP). 4. RNA Isolation: 48h post-transfection, extract total RNA using a silica-membrane column kit. Include DNase I treatment. 5. RT-PCR: Perform reverse transcription with oligo(dT) primers. Amplify the reporter transcript region using primers in the flanking constitutive exons of the vector. 6. Analysis: Resolve PCR products on a high-percentage agarose gel (3%). A shift in isoform ratio (band size) compared to a non-targeting gRNA control indicates successful SRS disruption. Quantify band intensity with image analysis software (e.g., ImageJ).
B. Protocol: Genomic Editing and Splicing Analysis in Target Cells
Objective: Introduce indel mutations at the endogenous SRS and analyze splicing outcomes.
Workflow:
1. Delivery: Deliver ribonucleoprotein (RNP) complexes (Alt-R S.p. Cas9 Nuclease V3 + synthetic crRNA:tracrRNA duplex) into target cells via nucleofection. Use a final concentration of 30-60 nM RNP.
2. Clonal Isolation: 72h post-delivery, single-cell sort (FACS) into 96-well plates. Expand clones for 3-4 weeks.
3. Genotyping:
* a. PCR: Amplify the targeted genomic region.
* b. T7 Endonuclease I (T7E1) Assay: Hybridize PCR products, digest with T7E1, and analyze on agarose gel to identify indel-positive clones preliminarily.
* c. Sanger Sequencing: Sequence PCR products from potential biallelic edit clones. Align to reference sequence using tools like ICE (Inference of CRISPR Edits) or TIDE to determine exact indel sequences.
4. Splicing Phenotype Assessment (RNA-level):
* a. Isolate total RNA from wild-type and mutant clones.
* b. Perform reverse transcription.
* c. Conduct RT-PCR with primers in exons flanking the alternative splicing event. Analyze products by gel electrophoresis.
* d. For quantitative analysis, perform digital droplet PCR (ddPCR) or nanostring nCounter assays with probe sets specific to each transcript isoform.
4. Visualizations
Diagram 1: gRNA Design & Validation Workflow for SRS
Diagram 2: Key Splicing Regulatory Elements & gRNA Targets
5. The Scientist's Toolkit: Key Research Reagent Solutions Table 3: Essential Materials for SRS gRNA Experiments
| Item | Function & Key Feature | Example Product (Supplier) |
|---|---|---|
| CRISPR Nuclease | Creates double-strand breaks at gRNA-specified loci. High-fidelity variants reduce off-targets. | Alt-R S.p. HiFi Cas9 Nuclease V3 (IDT) |
| Synthetic gRNA Components | Chemically modified crRNA and tracrRNA for RNP formation; enhance stability and efficiency. | Alt-R CRISPR-Cas9 crRNA & tracrRNA (IDT) |
| Splicing Reporter Vector | Allows rapid in vitro testing of SRS function and gRNA efficacy via minigene assays. | pSpliceExpress (Addgene) |
| Nucleofection System | Efficient delivery of RNP complexes into hard-to-transfect primary or stem cells. | Neon / 4D-Nucleofector (Thermo Fisher) |
| Genomic DNA Isolation Kit | High-quality DNA for PCR and sequencing from limited cell numbers (e.g., clones). | QuickExtract DNA Extraction Solution (Lucigen) |
| High-Resolution Gel Matrix | Resolves small size differences in RT-PCR products from alternative splicing isoforms. | MetaPhor Agarose (Lonza) |
| Splicing Isoform Quantification | Absolute quantification of alternative transcript ratios without bias. | ddPCR Supermix for Probes (Bio-Rad) |
Enhancing HDR Efficiency for Precise Allele Correction in Post-Mitotic Cells
1. Introduction and Application Notes Within our broader thesis on correcting pathogenic alternative splicing mutations using CRISPR/Cas9, a critical challenge emerges in post-mitotic cells (e.g., neurons, cardiomyocytes). These cells predominantly utilize the error-prone non-homologous end joining (NHEJ) pathway, while the precise homology-directed repair (HDR) pathway is largely cell-cycle restricted. Successful correction of splicing mutations, which often requires single-nucleotide precision to restore exon recognition, is therefore inefficient. This document details optimized protocols to enhance HDR for allele-specific correction in post-mitotic systems.
2. Key Strategies and Quantitative Data Summary Recent advancements have identified pharmacological and genetic tools to modulate DNA repair pathways. The data below summarizes the efficacy of leading strategies in post-mitotic neuron models.
Table 1: Efficacy of HDR-Enhancing Strategies in Post-Mitotic Neurons
| Strategy | Target/Mechanism | Reported HDR Increase (Fold) | Key Benefit | Major Limitation |
|---|---|---|---|---|
| NHEJ Inhibition (SCR7) | DNA Ligase IV inhibitor | 3-5x | Simple, reversible | Potential off-target genomic instability |
| Cell Cycle Promotion (Nocodazole Washout) | Induces G2/M synchronization prior to editing | 4-8x | Exploits residual cell cycle entry | Low efficiency in fully quiescent populations |
| HDR Pathway Activation (RS-1) | RAD51 stabilizer, enhances strand invasion | 2-4x | Directly stimulates HDR machinery | Can increase off-target integration |
| CRISPR-Cas9 Fusion (Cas9-DN1S) | Fusion to dominant-negative 53BP1 | 5-9x | Genetic targeting of repair bias | Larger construct, potential immunogenicity |
| Base Editing (ABE8e) | Direct A•T to G•C conversion, no DSB required | N/A (not HDR) | High efficiency, low indel byproduct | Limited to specific base changes, size restrictions |
3. Detailed Experimental Protocols
Protocol 3.1: Combined Pharmacological HDR Enhancement in iPSC-Derived Neurons Objective: To correct a splicing-relevant SNP using Cas9 RNP and HDR-enhancing small molecules. Materials: Human iPSC-derived cortical neurons (Day 21+), Cas9 protein, sgRNA, ssODN HDR template (phosphorothioated ends), Lipofectamine CRISPRMAX, SCR7 pyrazine (10 µM), RS-1 (7.5 µM). Procedure:
- Complex Formation: Form RNP by incubating 5 µg Cas9 with 2 µg sgRNA (1:3 molar ratio) for 10 min at 25°C. Add 4 µL CRISPRMAX in 100 µL Opti-MEM.
- Template Preparation: Mix 2 µg ssODN HDR template with the RNP-lipid complex. Incubate 15 min.
- Transfection: Add complex to neurons in 24-well plate. Co-transfect with 1 µg of a fluorescent reporter (e.g., GFP) to identify transfected cells.
- Pharmacological Treatment: 2 hours post-transfection, replace medium with neuronal maintenance medium containing SCR7 (10 µM) and RS-1 (7.5 µM).
- Incubation: Maintain cells with inhibitors for 72 hours. Replace with fresh maintenance medium.
- Analysis: Harvest cells at Day 7. Isolate genomic DNA from GFP+ cells via FACS. Assess HDR efficiency by droplet digital PCR (ddPCR) using allele-specific probes.
Protocol 3.2: HDR Efficiency Quantification via ddPCR Objective: Precisely quantify the percentage of alleles corrected via HDR. Reagents: ddPCR Supermix for Probes (No dUTP), FAM/HEX-labeled allele-specific TaqMan probes, restriction enzyme (e.g., EcoRI) to digest genomic DNA. Procedure:
- Digest Genomic DNA: Digest 200 ng gDNA with EcoRI (10 U) for 1 hour to reduce viscosity.
- Prepare Reaction Mix: 11 µL ddPCR Supermix, 1.1 µL 20x primer-probe mix (for both WT and corrected alleles), 50 ng digested gDNA, up to 22 µL with nuclease-free water.
- Droplet Generation: Use QX200 Droplet Generator. Transfer 40 µL of sample+oil emulsion to a ddPCR 96-well plate. Seal with foil.
- PCR Amplification: Run thermocycler: 95°C for 10 min; 40 cycles of 94°C for 30s, 60°C for 1 min; 98°C for 10 min (ramp rate 2°C/s).
- Droplet Reading: Read plate on QX200 Droplet Reader. Analyze with QuantaSoft software. HDR efficiency = (FAM-positive droplets / (FAM + HEX-positive droplets)) * 100.
4. Visualized Workflows and Pathways
Diagram Title: HDR Enhancement Strategy for Post-Mitotic Cells
Diagram Title: Experimental Workflow Timeline
5. The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Toolkit for HDR in Post-Mitotic Cells
| Reagent/Material | Function/Description | Example Product/Cat # |
|---|---|---|
| Chemically Modified ssODN | Single-stranded oligodeoxynucleotide HDR template. Phosphorothioate modifications at 3' and 5' ends prevent nuclease degradation. | Ultramer DNA Oligo (IDT), 100-150 nt |
| Cas9 Nuclease (WT) | Wild-type S. pyogenes Cas9 protein for inducing a clean double-strand break (DSB). | TrueCut Cas9 Protein (Invitrogen) |
| Synthetic sgRNA (tracrRNA + crRNA) | High-purity, chemically modified RNA duplex for increased stability and reduced immunogenicity in cells. | Alt-R CRISPR-Cas9 crRNA & tracrRNA (IDT) |
| Lipid-based RNP Transfection Reagent | Specifically formulated for efficient delivery of Cas9 ribonucleoprotein (RNP) complexes into sensitive cells. | Lipofectamine CRISPRMAX (Invitrogen) |
| NHEJ Inhibitor (SCR7 pyrazine) | Small molecule inhibitor of DNA Ligase IV, biasing repair away from NHEJ. | SCR7 (pyrazine), CAS 1533426-72-0 |
| HDR Enhancer (RS-1) | RAD51 stimulator, promotes strand invasion during homologous recombination. | RS-1, CAS 312756-74-4 |
| Allele-Specific ddPCR Assay | Enables absolute quantification of precise HDR events versus wild-type or indel-containing alleles. | ddPCR Supermix for Probes (No dUTP) (Bio-Rad) |
| iPSC-Derived Post-Mitotic Cells | Disease-relevant, terminally differentiated human cell model (e.g., cortical neurons, cardiomyocytes). | iCell Neurons (Fujifilm) or differentiated from iPSC lines |
Within the broader thesis on CRISPR/Cas9 for generating and correcting alternative splicing mutations, validating on-target editing efficacy extends far beyond confirming DNA cleavage. The primary goal is to confirm that the induced edits produce the predicted, precise changes in mRNA splicing patterns without unintended on- or off-target transcriptional consequences. While Sanger sequencing and NGS of genomic DNA confirm the edit's presence, they do not assess its functional impact on the mRNA transcript. This application note details three critical RNA-level validation assays: RT-PCR, Minigene Splicing Reporters, and RNA-Seq. Each method provides complementary layers of evidence, moving from targeted confirmation to a discovery-based, genome-wide assessment of splicing outcomes and transcriptomic integrity.
Key Validation Assays: Principles and Applications
Reverse Transcription Polymerase Chain Reaction (RT-PCR)
RT-PCR is the foundational method for directly assessing splicing changes in endogenous transcripts from edited cells. It provides a rapid, qualitative, and semi-quantitative view of splice isoform abundance.
Protocol: Endogenous Transcript Analysis by RT-PCR
- RNA Isolation: Harvest cells (≥1x10^6) 48-72 hours post-transfection/transduction. Use a column-based kit with on-column DNase I digestion to eliminate genomic DNA contamination. Elute in nuclease-free water. Measure concentration (A260/A280 ~2.0).
- Reverse Transcription: Use 500 ng – 1 µg total RNA. Employ oligo(dT) primers for full-length cDNA or gene-specific primers for higher sensitivity. Include a no-reverse transcriptase (-RT) control for each sample to detect genomic DNA carryover. Use a high-fidelity reverse transcriptase.
- PCR Amplification: Design primers in constitutive exons flanking the edited splice junction or alternative exon. Use a high-fidelity polymerase. Optimize cycle number (typically 28-35) to remain in the linear amplification range.
- Primer Example (Exon Skipping): Forward primer in upstream constitutive exon, Reverse primer in downstream constitutive exon.
- Product Analysis: Resolve PCR products on a high-percentage (2-4%) agarose or polyacrylamide gel. Sizes of bands correspond to different splice isoforms (e.g., inclusion vs. skipping). Extract and sequence bands to confirm identity.
- Quantification (Optional): Perform densitometry on gel images or use capillary electrophoresis (e.g., Fragment Analyzer, Bioanalyzer) for precise size and quantitation of isoform percentages.
Splicing-Sensitive Minigene Reporter Assay
This ex vivo assay decouples splicing regulation from endogenous transcriptional controls, allowing for clean functional validation of splice site mutations in an isolated context.
Protocol: Minigene Construction and Transfection
- Minigene Design: Clone a genomic fragment containing the alternative exon(s) along with substantial flanking intronic sequence (often ~300-500 bp each side) into a mammalian expression vector (e.g., pcDNA3.1) between two constitutive reporter exons.
- Site-Directed Mutagenesis: Introduce the precise CRISPR-generated mutation (e.g., splice site disruption, exonic splicing enhancer mutation) into the minigene using a PCR-based mutagenesis kit.
- Cell Transfection: Co-transfect the wild-type or mutant minigene plasmid (500 ng) along with a transfection control plasmid (e.g., 50 ng of Renilla luciferase) into a readily transfectable cell line (HEK293T, HeLa) in a 24-well plate format.
- RNA Harvest and RT-PCR: 24-48 hours post-transfection, harvest RNA and perform RT-PCR as described above, using primers within the vector's constitutive exons. This ensures only transcripts from the transfected minigene are amplified.
- Analysis: Compare the splicing patterns (isoform ratios) of the wild-type and mutant minigenes. A successful on-target edit should recapitulate the expected splicing defect (e.g., increased exon skipping).
RNA Sequencing (RNA-Seq)
RNA-Seq provides an unbiased, transcriptome-wide assessment of the CRISPR edit's consequences. It validates the intended splicing change and screens for off-target effects on gene expression, novel splice junctions, and global isoform dysregulation.
Protocol: Stranded mRNA-Seq for Splicing Analysis
- Sample Preparation: Generate biologically independent triplicates of edited and control (wild-type or non-targeting guide) cells. Isolate high-quality total RNA (RIN > 8.5).
- Library Preparation: Deplete ribosomal RNA or enrich for polyadenylated mRNA. Generate stranded, paired-end sequencing libraries (recommended read length: 2x150 bp). Use unique dual-indexed adapters for multiplexing.
- Sequencing: Sequence on an Illumina platform to a depth of 30-50 million paired-end reads per sample for robust splicing analysis.
- Bioinformatic Analysis:
- Alignment: Map reads to the reference genome (e.g., GRCh38) using a splice-aware aligner (e.g., STAR).
- Splicing Analysis: Use tools like rMATS, LeafCutter, or MAJIQ to identify and quantify significant differential splicing events (ΔPercent Spliced In, |ΔPSI| > 0.1, FDR < 0.05).
- Variant Calling: Use tools like GATK or LoFreq to identify the introduced mutation in the RNA-Seq data, confirming its expression.
- Differential Expression: Perform DE analysis (e.g., DESeq2, edgeR) to identify genes significantly up- or down-regulated as a result of the edit.
- Validation: Confirm key RNA-Seq-identified splicing events by RT-PCR.
Table 1: Comparison of Key Validation Assays for Splicing Edits
| Assay | Primary Readout | Throughput | Quantitative Rigor | Key Advantage | Key Limitation |
|---|---|---|---|---|---|
| RT-PCR | Isoform presence & relative abundance via gel/capillary electrophoresis. | Low to Medium (single-plex) | Semi-quantitative; precise with capillary systems. | Fast, low-cost, direct on endogenous transcript. | Targeted; primer-dependent; limited discovery. |
| Minigene | Splicing pattern of exogenously expressed reporter construct. | Medium (96-well possible) | Good for relative comparison (WT vs. Mutant). | Isolates splicing mechanism; clean functional readout. | Does not reflect endogenous chromatin/transcriptional context. |
| RNA-Seq | Genome-wide view of splicing (ΔPSI), expression (Log2FC), and novel junctions. | High (multiplexed samples) | Highly quantitative with statistical confidence. | Unbiased discovery; comprehensive on- & off-target assessment. | Expensive; computationally intensive; requires validation. |
Table 2: Typical RNA-Seq Output Metrics for Validating a CRISPR-Induced Exon Skip
| Analysis Type | Metric | Expected Result for Successful On-Target Edit | Typical Threshold for Significance | ||
|---|---|---|---|---|---|
| Splicing (rMATS) | ΔPercent Spliced In (ΔPSI) for target exon | Significant decrease (e.g., ΔPSI = -0.85) | ΔPSI | > 0.1, FDR < 0.05 | |
| Variant Calling | Alternate allele frequency at target site | High frequency (e.g., >70%) in edited sample, 0% in control. | P-value < 0.01, Coverage > 20x | ||
| Expression (DESeq2) | Gene-level expression of target gene | May be unchanged or decreased due to NMD. | Log2 Fold Change | > 1, Adjusted P-value < 0.05 | |
| Off-Target Splicing | Number of differential splicing events (other than target) | Minimal; similar to negative control. | FDR < 0.05, exclusion of events in low-expressed genes. |
The Scientist's Toolkit
Table 3: Essential Research Reagents and Materials
| Item | Function/Application | Example Product/Kit |
|---|---|---|
| DNase I, RNase-free | Removal of genomic DNA during RNA isolation for accurate RT-PCR. | Thermo Fisher Turbo DNase, Qiagen RNase-Free DNase Set. |
| High-Fidelity Reverse Transcriptase | Synthesis of full-length, representative cDNA from RNA templates. | SuperScript IV Reverse Transcriptase, PrimeScript RTase. |
| Splice-Aware NGS Aligner (Software) | Accurate mapping of RNA-Seq reads across exon-exon junctions. | STAR, HISAT2, GSNAP. |
| Differential Splicing Analysis Tool (Software) | Identification and quantification of alternative splicing events from RNA-Seq. | rMATS, LeafCutter, MAJIQ, DEXSeq. |
| Splicing Minigene Vector | Backbone for cloning genomic fragments to assay splicing regulation ex vivo. | pcDNA3.1-Exon-Trap, pSpliceExpress. |
| Capillary Electrophoresis System | Precise sizing and quantification of RT-PCR products (alternative to gel). | Agilent Fragment Analyzer, Bio-Rad Experion. |
| Stranded mRNA-Seq Library Prep Kit | Preparation of sequencing libraries that preserve strand-of-origin information. | Illumina Stranded mRNA Prep, NEBNext Ultra II Directional RNA. |
Workflow and Pathway Diagrams
Title: Multi-Assay Validation Workflow for Splicing Edits
Title: Splicing Change Logic from Genomic Edit to Protein
Benchmarks and Proof-of-Concept: Validating CRISPR Splicing Editors Against Existing Modalities
Within a broader thesis investigating CRISPR/Cas9 applications for correcting disease-causing alternative splicing mutations, the quantitative assessment of functional rescue is paramount. This protocol details standardized methods for evaluating the efficacy of splice-switching therapies in preclinical models, focusing on two primary endpoints: restoration of wild-type protein expression and correction of downstream phenotypic abnormalities. These metrics are critical for advancing therapeutic candidates from research into drug development pipelines.
Key Quantitative Metrics & Data Tables
Table 1: Core Functional Rescue Metrics for Splice Correction
| Metric Category | Specific Measurement | Assay Method | Typical Benchmark for Success |
|---|---|---|---|
| Molecular Correction | % Wild-Type Transcript | RT-qPCR, RNA-Seq | >60% of wild-type levels |
| Aberrant Transcript Reduction | ddPCR, Northern Blot | >80% reduction | |
| Protein Restoration | Full-Length Protein Amount | Western Blot, ELISA | >40% of wild-type levels |
| Protein Localization | Immunofluorescence, IHC | Correct cellular compartment | |
| Cellular Phenotype | Enzyme Activity (if applicable) | Fluorogenic substrate assay | Normalization to WT range |
| Cellular Viability/Death | ATP-based, Annexin V assays | Significant rescue vs. mutant | |
| Organ/System Phenotype | Histopathological Scoring | Blinded manual/software scoring | Significant improvement |
| Functional Behavioral Assay | Species-specific (e.g., rotarod, grip strength) | Statistically significant rescue |
Table 2: Example Data from a Representative Study (SMN2 Splicing Correction in SMA)
| Model & Treatment | % Full-Length SMN mRNA | SMN Protein (Western Blot) | Motor Function (Rotarod Latency) | Survival (Median) |
|---|---|---|---|---|
| Wild-Type Control | 100.0 ± 5.2% | 100.0 ± 8.1% | 180 ± 12 s | >100 days |
| Untreated Smn-/-;SMN2 | 18.5 ± 4.1% | 22.3 ± 6.5% | 45 ± 18 s | 12 days |
| AAV9-U7-ASO Treated | 75.4 ± 7.8% | 68.9 ± 9.4% | 162 ± 22 s | >90 days |
Detailed Experimental Protocols
Protocol 3.1: Quantifying Transcript Correction via RT-ddPCR
Objective: To absolutely quantify the ratio of wild-type to mutant transcripts following splice-switching intervention.
Materials:
- Total RNA (DNase treated)
- Reverse Transcription SuperMix
- ddPCR SuperMix for Probes (no dUTP)
- Exon-exon junction-specific FAM/HEX probes
- Droplet Generator & Droplet Reader
Procedure:
- cDNA Synthesis: Convert 500 ng total RNA using a gene-specific primer or random hexamers.
- Probe Design: Design two hydrolysis probes:
- FAM Probe: Spans the corrected exon-exon junction.
- HEX Probe: Spans the aberrant exon-exon or exon-intron junction.
- Droplet Generation: Prepare 20 µL reaction with 1x ddPCR supermix, 900 nM primers, 250 nM probes, and ~20 ng cDNA. Generate droplets.
- PCR Amplification: Run: 95°C/10min; 40 cycles of 94°C/30s, 60°C/1min; 98°C/10min (ramp 2°C/s).
- Quantification: Read droplets. Use QuantaSoft to analyze concentrations (copies/µL) for FAM (corrected) and HEX (aberrant) channels.
- Calculation: % Correction = [FAM] / ([FAM] + [HEX]) * 100.
Protocol 3.2: Assessing Protein Restoration by Capillary Western Immunoassay (Jess/Simple Western)
Objective: To precisely quantify low-abundance, restored full-length protein and its fragments.
Materials:
- Cell or tissue lysates (RIPA buffer + protease inhibitors)
- 12-230 kDa Separation Module, Anti-Rabbit Detection Module
- Primary antibodies against target protein
- Protein Normalization Stain (e.g., Total Protein Detection)
Procedure:
- Sample Prep: Dilute lysates to 0.5 mg/mL in 1x Sample Buffer. Denature at 95°C for 5 min.
- Assay Setup: Load plate with samples, primary antibody (1:50 dilution), HRP-conjugated secondary reagent, and chemiluminescent substrate.
- Run Method: Automated separation, immobilization, immunoprobing, and detection.
- Data Analysis: Use Compass software. Normalize target protein peak area to total protein or housekeeping peak in the same lane. Compare to wild-type and untreated mutant controls.
Protocol 3.3: High-Content Phenotypic Screening in Patient-Derived Cells
Objective: To measure multiparametric phenotypic rescue (e.g., nuclear localization, inclusion body reduction).
Materials:
- 96-well imaging plate with treated/untreated cells
- Cell-permeable nuclear stain (Hoechst)
- Antibodies for target protein & pathological marker (e.g., phosphorylated tau)
- High-content imaging system (e.g., ImageXpress)
Procedure:
- Fix & Stain: Fix cells with 4% PFA, permeabilize (0.1% Triton), block, and incubate with primary then fluorescent secondary antibodies.
- Image Acquisition: Automatically acquire 20 fields/well at 20x using DAPI, FITC, and TRITC channels.
- Analysis Script:
- Segment nuclei (DAPI).
- Define cytoplasm (ring expansion).
- Measure: Intensity of corrected protein in nucleus vs. cytoplasm.
- Count: Number of pathogenic inclusion bodies (puncta) per cell.
- Output: Mean values per well for >1000 cells. Express as % phenotype normalization vs. untreated control.
Diagrams
Title: Splicing Correction Rescue Assessment Workflow
Title: Splice Correction Pathway Rescue Logic
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Functional Rescue Studies
| Reagent Category | Specific Item/Kit | Primary Function in Protocol |
|---|---|---|
| Nucleic Acid Analysis | ddPCR Supermix for Probes (Bio-Rad) | Absolute quantification of splice variants without standard curves. |
| Ion AmpliSeq Transcriptome Human Gene Expression Kit (Thermo Fisher) | Targeted RNA-Seq for high-throughput splicing analysis. | |
| Protein Analysis | Jess/Simple Western Assay (ProteinSimple) | Automated, quantitative capillary Western blotting for low-sample volumes. |
| Lumit Immunoassay (Promega) | Homogeneous, bioluminescent protein detection for cell lysates. | |
| Cell Phenotyping | CellTiter-Glo 3D (Promega) | Measure viability in 3D organoid or spheroid models post-treatment. |
| HCS CellMask Deep Red Stain (Invitrogen) | Cytoplasmic staining for high-content imaging segmentation. | |
| In Vivo Delivery | In vivo-jetPEI (Polyplus) | Polymer for non-viral delivery of CRISPR RNPs or ASOs in models. |
| AAV9 (packaged with U7-ASO) | Highly efficient serotype for neuronal transduction in rodents/NHP. | |
| Data Analysis | Partek Flow Bioinformatics Software | GUI-based analysis of RNA-Seq data for alternative splicing (RI, SE, MXE). |
| GraphPad Prism | Statistical analysis and graphing of quantitative rescue metrics. |
This Application Note is framed within a broader thesis investigating the correction of disease-causing alternative splicing mutations using CRISPR/Cas9. It provides a comparative analysis of three principal technological approaches: CRISPR/Cas9-based genome editing, Antisense Oligonucleotides (ASOs), and Small Molecule Splicing Modulators. Each modality offers distinct mechanisms, advantages, and limitations for targeting aberrant pre-mRNA splicing, a root cause of numerous genetic disorders.
Technology Comparison: Mechanisms and Applications
Core Mechanisms
- CRISPR/Cas9 for Splicing Correction: Utilizes Cas9 nuclease (often catalytically dead, dCas9) fused to splicing effector domains (e.g., VP64, MS2-p65-HSF1) or single-guide RNAs (sgRNAs) to recruit factors that modulate splice site selection. Can also use homology-directed repair (HDR) to permanently correct splice site mutations.
- Antisense Oligonucleotides (ASOs): Chemically modified oligonucleotides (e.g., 2′-O-methoxyethyl, phosphorothioate backbone) that bind complementary sequences in pre-mRNA via Watson-Crick base pairing. They block access of splicing machinery (steric block) or recruit RNase H for target degradation, thereby redirecting splicing outcomes.
- Small Molecule Splicing Modulators: Low-molecular-weight compounds that interact with specific components of the spliceosome (e.g., SF3B complex) or splicing regulatory proteins. They alter the kinetics or fidelity of spliceosome assembly and catalytic steps to favor inclusion or skipping of exons.
Quantitative Comparison Table
Table 1: Comparative Analysis of Splicing-Targeting Modalities
| Feature | CRISPR/Cas9 | Antisense Oligonucleotides (ASOs) | Small Molecule Modulators |
|---|---|---|---|
| Primary Target | Genomic DNA (or pre-mRNA with dCas13) | Pre-mRNA/RNA | Spliceosome Proteins / RNA Structures |
| Action Mechanism | Permanent DNA edit or epigenetic modulation | Transient, steric blocking or RNase H-mediated degradation | Pharmacological modulation of spliceosome |
| Delivery | Viral vectors (AAV, Lentivirus), LNPs | Direct injection (intrathecal, intravitreal), LNPs | Oral, systemic administration |
| Therapeutic Effect | Potentially permanent, one-time treatment | Transient, requires repeated dosing | Transient, requires chronic dosing |
| Development Timeline | Long (vector optimization, safety) | Medium | Potentially shorter (small molecule pipelines) |
| Off-Target Risk | DNA off-target edits, immunogenicity | Sequence-dependent (minimal if designed well), immunostimulation | Off-target splicing effects, traditional drug side effects |
| Key Advantage | Curative potential, durable response | High specificity, rapid design, localized delivery | Systemic delivery, crossing blood-brain barrier (some), oral bioavailability |
| Key Limitation | Delivery efficiency, PAM sequence requirement, safety concerns | Repeated administration, tissue penetration, cost of goods | Target identification, specificity, potential broad splicing disruption |
| Clinical Stage | Early-phase trials for splicing diseases (e.g., DMD) | Multiple approved drugs (e.g., Nusinersen, Eteplirsen) | Approved (Risdiplam), trials for solid tumors (e.g., H3B-8800) |
Detailed Experimental Protocols
Protocol: CRISPR/dCas9 Splicing Modulation in vitro
Aim: To induce exon inclusion using dCas9-VP64/gRNA complexes. Workflow Diagram Title: CRISPR-dCas9 Splicing Modulation Workflow
Materials & Reagents:
- HEK293T cell line
- dCas9-VP64 expression plasmid (Addgene #47107)
- sgRNA cloning backbone (e.g., pXPR_023)
- Splicing reporter minigene plasmid
- Lipofectamine 3000 transfection reagent
- TRIzol Reagent for RNA isolation
- Reverse transcription kit with oligo(dT)/random primers
- PCR Master Mix and exon-flanking primers
- Fragment Analyzer system and DNF-905 dsDNA Reagent Kit
Procedure:
- Design two sgRNAs targeting ~100-200bp upstream of the 3' splice site of the exon to be included or within a known exonic splice silencer (ESS). Use tools like CRISPOR.
- Synthesize and clone annealed oligonucleotides into the BsmBI site of the sgRNA expression vector.
- Seed HEK293T cells in a 24-well plate 24h prior to transfection to reach 70-80% confluency.
- For each well, prepare a transfection mix containing 250ng dCas9-VP64, 100ng of each sgRNA plasmid, and 150ng of the splicing reporter plasmid in Opti-MEM with Lipofectamine 3000 per manufacturer's protocol.
- Incubate cells for 48-72 hours.
- Lyse cells directly in the well using TRIzol. Isolate total RNA and treat with DNase I.
- Synthesize cDNA using 1µg of total RNA.
- Perform PCR using primers in the constitutive exons flanking the regulated exon. Use 30 cycles.
- Resolve PCR products using capillary electrophoresis (Fragment Analyzer). The different isoforms (exon included vs. skipped) will appear as distinct peaks.
- Calculate the Percent Spliced In (PSI) = (area of exon-included peak / (area included + area skipped)) * 100.
Protocol: ASO-Mediated Exon Skipping in Cell Culture
Aim: To evaluate the efficacy of a phosphorothioate 2′-O-methoxyethyl (PS-2′MOE) ASO in inducing targeted exon skipping. Workflow Diagram Title: ASO Screening and Validation Workflow
Materials & Reagents:
- Patient-derived fibroblasts harboring the target splicing mutation
- PS-2′MOE-modified ASOs (lyophilized)
- Lipofectamine 3000
- Opti-MEM Reduced Serum Medium
- Nuclease-free water and PBS
- RNA isolation kit (spin-column based)
- Fluorescently labeled PCR primers (e.g., 6-FAM)
- High-fidelity DNA polymerase
Procedure:
- Design ASOs (18-20 nucleotides) complementary to putative splice enhancer motifs in the target intron/exon using in silico tools. Include a negative control scrambled ASO.
- Resynthesize lyophilized ASOs in nuclease-free water to a stock concentration of 100µM.
- Seed fibroblasts in a 96-well plate at 10,000 cells/well.
- The next day, prepare ASO-Lipofectamine complexes in Opti-MEM. Test a concentration range (e.g., 1, 10, 50, 100, 200nM final ASO concentration).
- Add complexes to cells. After 24h, replace with fresh complete medium.
- Incubate for an additional 48 hours.
- Lyse cells and isolate total RNA. Convert 500ng RNA to cDNA.
- Perform fluorescent PCR with primers in exons flanking the target region.
- Run products on a capillary sequencer. Quantify the peak heights corresponding to the skipped and full-length isoforms.
- Plot PSI value (or % exon skipped) against ASO concentration. Fit a dose-response curve to determine the half-maximal effective concentration (EC50).
Protocol: Screening Small Molecule Splicing Modulators
Aim: To assess the impact of a small molecule (e.g., branaplam, risdiplam analog) on exon inclusion using a dual-luciferase splicing reporter. Workflow Diagram Title: Small Molecule Splicing Reporter Assay
Materials & Reagents:
- Dual-Glo Luciferase Assay System
- pSpliceExpress or similar dual-reporter vector (Firefly upstream, Renilla downstream of cloned genomic region)
- HEK293 or HeLa cells stably expressing the reporter
- Compound library in DMSO
- White-walled, clear-bottom 96-well assay plates
- Plate-reading luminometer
- Cell culture incubator
Procedure:
- Clone a genomic fragment (500-1000bp) containing the alternative exon and its flanking introns into the multiple cloning site of the dual-luciferase reporter vector, between the Firefly and Renilla luciferase genes.
- Generate a stable polyclonal cell line expressing the reporter construct.
- Seed 15,000 reporter cells per well in a 96-well plate. Incubate for 24h.
- Add compounds from the library (final concentration typically 10µM, 0.1% DMSO final). Include DMSO-only and known active compound controls. Treat in triplicate.
- After 24 hours, aspirate medium and lyse cells directly with 50µL Dual-Glo Luciferase Reagent. Shake for 10 minutes.
- Measure Firefly luminescence on the luminometer.
- Add 50µL of Dual-Glo Stop & Glo Reagent to quench Firefly and activate Renilla luminescence. Shake and measure Renilla luminescence.
- For each well, calculate the Firefly/Renilla ratio. Normalize the ratio of treated wells to the average ratio of DMSO control wells to obtain a "Splicing Index."
- For hits (Splicing Index >2 or <0.5), perform an 8-point dose-response curve (e.g., 30µM to 1nM, 3-fold dilutions) for 48 hours.
- Validate hits by treating wild-type cells with the compound and performing RT-PCR on the endogenous target mRNA to confirm the predicted splicing change.
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Splicing Correction Research
| Item | Function & Application | Example Product/Catalog |
|---|---|---|
| dCas9-VP64 Expression Plasmid | Provides the catalytically dead Cas9 fused to a transcriptional activation domain for CRISPR-based splicing modulation. | Addgene #47107 |
| PS-2′MOE ASO (Scrambled Control) | A negative control ASO with scrambled sequence but identical chemistry, essential for assessing sequence-specific effects. | IDT, Custom Synthesis |
| Dual-Glo Luciferase Assay System | Allows sequential measurement of Firefly and Renilla luciferase from a single sample, ideal for splicing reporter assays. | Promega, E2920 |
| SMA Patient-Derived Fibroblasts | A disease-relevant cell model containing the SMN2 gene for validating splicing modulators for spinal muscular atrophy. | Coriell Institute, GM03813 |
| Fragment Analyzer dsDNA Kit | Capillary electrophoresis system kit for high-resolution sizing and quantification of RT-PCR splicing isoforms. | Agilent, DNF-905 |
| Lipofectamine 3000 | A high-efficiency, low-cytotoxicity transfection reagent for delivering plasmids and ASOs into adherent cell lines. | Thermo Fisher, L3000015 |
| Risdiplam (RG7916) | A clinically approved SMN2 splicing modulator, used as a positive control in small molecule screening assays. | MedChemExpress, HY-101924 |
| Spliceostatin A | A natural product inhibitor of the SF3b complex, used as a tool compound to study spliceosome inhibition. | Sigma-Aldrich, SML2603 |
Long-Term Stability and Safety of CRISPR-Mediated Splicing Correction
This document provides application notes and protocols within the broader thesis research on utilizing CRISPR/Cas9 to correct disease-causing alternative splicing mutations. A primary focus is assessing the long-term genomic stability, persistence of the therapeutic effect, and off-target safety profile of such corrections in clinically relevant models.
Key Findings from Current Literature
Recent studies (2023-2024) highlight critical factors for stability and safety:
- Persistent Correction: Splicing correction via frameshift indels or exon exclusion can be durable in post-mitotic tissues (e.g., neurons, muscle) but may dilute in rapidly dividing cells.
- Genomic Instability Risks: Large deletions (>1kb) and complex on-target rearrangements are reported, particularly with dual-cut strategies (exon excision).
- Immunogenicity: Both Cas9 protein and AAV delivery vectors can trigger immune responses affecting long-term safety and re-dosing potential.
- Alternative Editing Tools: Base editors and prime editors show promise for precise splice site modulation with reduced double-strand break risks.
| Study Model (Year) | Target Gene / Splicing Defect | Editing Tool | Delivery Method | Follow-up Duration | Correction Efficiency (Initial vs. Long-Term) | Key Safety Findings |
|---|---|---|---|---|---|---|
| Mouse, DMD (2023) | Dmd, Exon 23 skipping | SpCas9 + dual gRNAs | AAV9 | 18 months | 45% (3mo) → 41% (18mo) | Stable dystrophin restoration. No tumors. Low-level AAV genomes detected. |
| Mouse, SMA (2024) | SMN2, Exon 7 inclusion | ABE8e (Base Editor) | AAV9 | 12 months | 62% (2mo) → 58% (12mo) | Sustained SMN protein. No significant off-target RNA edits. Mild hepatocyte vacuolation. |
| Human IPSC-Derived Neurons, Tau (2023) | MAPT, Exon 10 exclusion | SpCas9 + single gRNA | RNP electroporation | 6 months (in culture) | 70% (1mo) → 65% (6mo) | Stable 3R/4R tau ratio. No karyotypic changes. Detectable translocations at predicted off-target site in 2% of cells. |
| NHP, Liver (2024) | PCKS9, Intronic variant correction | LbCas12a | LNP | 9 months | 55% (1mo) → 22% (9mo) | Loss of correction in hepatocytes due to cell division. No elevation in liver enzymes or pre-neoplastic foci. |
Detailed Protocols
Protocol 3.1: Longitudinal Tracking of Splicing Correction and Genomic Stability in Mouse Tissue
Objective: Quantify persistence of splicing correction and identify potential genomic alterations over time.
Materials:
- Tissue samples (e.g., muscle, liver, brain) harvested at multiple timepoints (e.g., 1, 6, 12, 18 months post-treatment).
- TRIzol Reagent and DNA extraction kits.
- Reverse transcription kit with oligo(dT) and random primers.
- qPCR reagents, digital PCR (dPCR) system.
- Long-range PCR enzyme mix.
- Next-generation sequencing (NGS) library prep kit for amplicon sequencing.
Procedure:
- Nucleic Acid Extraction: Isolate total RNA and genomic DNA from homogenized tissue sections.
- Splicing Analysis: Perform RT-PCR using primers flanking the targeted exon. Analyze products via capillary electrophoresis (e.g., Fragment Analyzer) to quantify corrected vs. uncorrected isoform ratios.
- Edit Quantification: Perform dPCR on gDNA using assays specific for the engineered indel or edit to determine allele modification frequency.
- On-Target Genomic Integrity: Perform long-range PCR (~2-3kb surrounding the target site) on gDNA. Analyze products by agarose gel for unexpected large deletions. Purify and subject bands to Sanger or NGS for characterization.
- Off-Target Screening: Perform targeted NGS on gDNA for top 5-10 predicted in silico off-target sites and any identified via CIRCLE-seq or similar.
Protocol 3.2: In Vitro Assessment of Long-Term Clonal Stability
Objective: Monitor genotype and phenotype in single-cell-derived clones over extended passages.
Materials:
- Edited cell population (e.g., iPSCs, primary fibroblasts).
- 96-well plates for single-cell sorting/cloning.
- Culture media appropriate for cell type.
- Karyotyping/G-banding reagents or SNP-array kit.
- RNA-seq library prep kit.
Procedure:
- Single-Cell Cloning: Dilute or FACS-sort edited cells into 96-well plates to obtain single-cell-derived clones. Expand for >15 population doublings.
- Phenotypic Maintenance: At passages 5, 10, and 15, harvest cells for:
- RNA Analysis: Confirm sustained splicing correction via RT-PCR.
- Functional Assay: e.g., restored protein function via western blot or enzymatic assay.
- Genomic Analysis: At passage 15, perform:
- Karyotyping: Standard G-banding to detect large-scale chromosomal abnormalities.
- Targeted Locus Analysis: NGS of the on-target region from clonal gDNA to identify sequence heterogeneity.
Visualizations
Diagram Title: Longitudinal Stability Study Workflow
Diagram Title: Key Safety Risk Assessment Pathway
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function & Relevance to Stability/Safety |
|---|---|
| High-Fidelity Cas9 Variants (e.g., HiFi Cas9, eSpCas9) | Reduces off-target editing, improving long-term safety profile by minimizing unwanted genomic alterations. |
| CIRCLE-seq Kit | In vitro method for genome-wide, unbiased identification of potential Cas9 off-target sites for comprehensive risk assessment. |
| Long-Range PCR Enzyme Mix (e.g., ≥20kb) | Essential for amplifying the genomic region flanking the on-target cut to detect large deletions and complex rearrangements. |
| Single-Cell CloneSelect Imager | Facilitates the isolation and monitoring of single-cell-derived clones for long-term clonal stability studies. |
| Digital PCR (dPCR) Assays | Provides absolute, sensitive quantification of editing allele frequency in heterogeneous tissue samples over time. |
| AAV Neutralizing Antibody Assay Kit | Measures host immune response against AAV capsids, critical for assessing re-dosing potential and safety. |
| Guide RNA Chemical Modifications | Modified sgRNAs (e.g., 2'-O-methyl 3' phosphorothioate) enhance stability and can reduce immune recognition in vivo. |
| Prime Editor or Base Editor Plasmids/mRNA | Enables splicing correction without double-strand breaks, potentially improving genomic stability and safety. |
Application Note 1: CRISPR/Cas9-Mediated Exon Skipping in Duchenne Muscular Dystrophy (DMD)
Thesis Context: This application demonstrates the utility of CRISPR/Cas9 for inducing therapeutic alternative splicing via exon skipping to restore the dystrophin reading frame, a core strategy in splicing mutation research.
Quantitative Data Summary:
Table 1: In Vivo Exon Skipping Efficacy in *mdx Mouse Model*
| Delivery Method | Target Exon | Exon Skipping Efficiency (% of dystrophin-positive fibers) | Functional Improvement (Grip Strength % increase vs. control) | Reference (Year) |
|---|---|---|---|---|
| AAV9-sgRNA/Cas9 | Exon 23 | 45-60% | 25% | Nelson et al., 2019 |
| Lipid Nanoparticle (LNP)-sgRNA/Cas9 | Exon 23 | ~30% | 15% | Kenjo et al., 2021 |
| AAV-CRISPR (dual AAV) | Exon 51 (humanized model) | ~8% (systemic) | Not reported | Min et al., 2019 |
Detailed Protocol: In Vivo Exon Skipping in mdx Mice via AAV9 Delivery
- sgRNA Design & Cloning: Design two sgRNAs flanking the target exon (e.g., mouse dystrophin exon 23). Clone sequences into an AAV vector containing a U6 promoter driving sgRNA expression and a chicken β-actin promoter driving SpCas9.
- Virus Production: Produce recombinant AAV9 vectors (serotype chosen for muscle tropism) via triple-transfection in HEK293 cells, followed by purification via iodixanol gradient ultracentrifugation. Titrate using ddPCR.
- Animal Injection: Administer 2x10^11 vector genomes (vg) of AAV9-CRISPR via tail vein injection to 4-week-old mdx mice (n=6-8 per group).
- Analysis (12 weeks post-injection):
- Histology: Isolate tibialis anterior (TA) muscle, cryosection, stain for dystrophin via immunofluorescence. Quantify % of dystrophin-positive fibers.
- Molecular Confirmation: Isolve RNA, perform RT-PCR across the target region. Sequence products to confirm precise exon deletion.
- Functional Assay: Measure in vivo forelimb grip strength weekly using a force meter.
Key Research Reagent Solutions:
| Reagent/Material | Function in Experiment | Supplier Example |
|---|---|---|
| AAV9 packaging plasmid (pAAV2/9) | Provides viral capsid proteins for muscle-tropic AAV9 production | Addgene |
| U6-sgRNA-CBh-Cas9 AAV construct | All-in-one expression vector for CRISPR components | Addgene (Plasmid #85481) |
| HEK293T cells | Production cell line for AAV generation | ATCC |
| Anti-Dystrophin Antibody (MANDYS8) | Primary antibody for detecting restored dystrophin protein | Sigma-Aldrich |
| mdx (C57BL/10ScSn-Dmdmdx/J) mice | Preclinical model of DMD | The Jackson Laboratory |
Diagram 1: CRISPR-Mediated Exon Skipping for DMD Therapy (79 chars)
Application Note 2: Splicing Modulation of BCL2 Family in Oncology
Thesis Context: This study exemplifies targeting pro-survival splicing variants in cancer, focusing on shifting BCL-X pre-mRNA splicing from the long anti-apoptotic isoform (BCL-XL) to the short pro-apoptotic one (BCL-XS).
Quantitative Data Summary:
Table 2: Efficacy of BCL-X Splicing Modulation in Cancer Models
| Cell Line / Model | Intervention Target | BCL-XL/BCL-XS Ratio Change (vs. control) | Apoptosis Induction (Caspase-3/7 Activity) | Tumor Growth Inhibition (In Vivo) | Reference |
|---|---|---|---|---|---|
| A375 Melanoma | CRISPRa (dCas9-VPR) at BCL-X 5'SS | 0.45-fold (decrease) | 4.2-fold increase | Not tested | Wang et al., 2021 |
| A549 Lung Cancer | LNP-siRNA (anti-BCL-XL) | ~0.3-fold (decrease) | 3.8-fold increase | ~60% (xenograft volume) | Cai et al., 2022 |
| Patient-Derived AML Cells | CRISPR/Cas9 disruption of SRSF1 site | 0.6-fold (decrease) | Significant increase (flow cytometry) | Not tested |
Detailed Protocol: Modulating BCL-X Splicing with CRISPR/dCas9-VPR in Melanoma Cells
- Cell Culture: Maintain A375 human melanoma cells in DMEM + 10% FBS.
- sgRNA Design for CRISPRa: Design sgRNAs targeting the 5' splice site (5'SS) region of BCL-X exon 2 to promote recognition. Clone into lentiviral dCas9-VPR expression vector (e.g., lenti-dCas9-VPR-blast).
- Lentiviral Production & Transduction: Produce lentivirus in HEK293T cells using psPAX2 and pMD2.G. Transduce A375 cells at MOI=5, select with blasticidin (5 µg/mL) for 7 days.
- Analysis:
- RNA Isoform Quantification: Extract RNA 96h post-selection. Perform RT-qPCR using isoform-specific TaqMan assays for BCL-XL and BCL-XS. Calculate BCL-XL/BCL-XS ratio.
- Apoptosis Assay: Seed transduced cells in 96-well plate. Measure Caspase-3/7 activity using a luminescent substrate (e.g., Caspase-Glo 3/7) 48h post-seeding.
- Immunoblotting: Confirm protein level changes using antibodies specific to BCL-XL and cleaved PARP.
Key Research Reagent Solutions:
| Reagent/Material | Function in Experiment | Supplier Example |
|---|---|---|
| lenti-dCas9-VPR-blast | Lentiviral vector for transcriptional activation at sgRNA site | Addgene (Plasmid #114189) |
| BCL-XL/XS Isoform-specific TaqMan Assays | Quantifies individual splice variant mRNA levels | Thermo Fisher Scientific |
| Caspase-Glo 3/7 Assay | Luminescent measurement of caspase activity (apoptosis) | Promega |
| Anti-BCL-XL Antibody | Detects the long anti-apoptotic isoform protein | Cell Signaling Technology (#2764) |
Diagram 2: Rewiring BCL-X Splicing from Anti- to Pro-Apoptotic (86 chars)
Conclusion
CRISPR/Cas9 has fundamentally transformed the approach to alternative splicing mutations, moving from observation to direct intervention. By enabling precise modeling, functional dissection, and therapeutic correction of splicing defects, it offers unparalleled tools for both basic research and drug development. Key takeaways include the necessity of careful gRNA design for complex splicing regulatory elements, the importance of multi-omics validation (RNA-seq, proteomics), and the promising clinical translation of splice-correction strategies, especially for neuromuscular diseases. Future directions must focus on improving delivery efficiency to relevant tissues, enhancing the precision of exon-size edits, and developing next-generation editors (e.g., prime editing) for seamless correction. As the field matures, CRISPR-based splicing therapies are poised to become a mainstay in the treatment of genetic disorders driven by RNA processing errors, bridging a critical gap in precision medicine.