CRISPR-Select vs. Limiting Dilution Cloning: A Definitive Guide for Cell Line Development Researchers
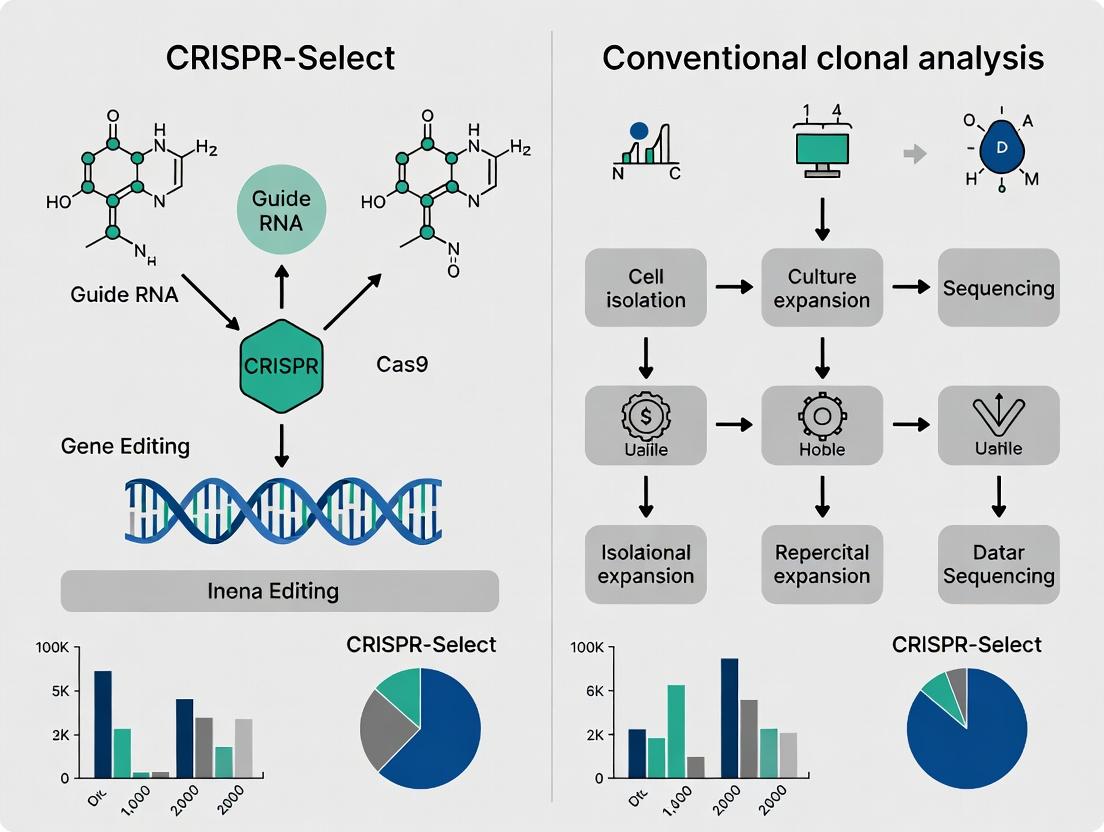
This article provides a comprehensive comparison for researchers, scientists, and drug development professionals between the novel CRISPR-Select workflow and traditional clonal analysis methods like limiting dilution.
CRISPR-Select vs. Limiting Dilution Cloning: A Definitive Guide for Cell Line Development Researchers
Abstract
This article provides a comprehensive comparison for researchers, scientists, and drug development professionals between the novel CRISPR-Select workflow and traditional clonal analysis methods like limiting dilution. We will explore the foundational principles of single-cell cloning, detail the step-by-step methodology of CRISPR-Select, address common troubleshooting and optimization challenges, and provide a rigorous validation and comparative analysis of throughput, monoclonality assurance, genotypic-phenotypic correlation, and cost-effectiveness. The goal is to equip the reader with the knowledge to select the optimal method for their specific gene-editing and cell line development projects.
The Essential Primer: Understanding the Need for Clonal Analysis in CRISPR-Cas9 Workflows
In the pursuit of understanding gene function, the generation of precisely engineered cell lines is foundational. While CRISPR-Cas9 enables targeted genomic modifications, the outcome is inherently heterogeneous. This comparison guide evaluates the paradigm of post-editing clonal isolation, contrasting traditional limiting dilution methods with the CRISPR-Select platform, to underscore why single-cell cloning is an indispensable step in functional genomics research.
Comparative Analysis: Clonal Isolation Methodologies
| Parameter | Traditional Limiting Dilution | CRISPR-Select Platform | Supporting Experimental Data |
|---|---|---|---|
| Workflow Duration | 4-6 weeks (including expansion & screening) | 2-3 weeks (integrated screening) | NGS validation shows CRISPR-Select identifies clones 14 days post-transfection vs. 28+ days for dilution. |
| Clonal Purity Assurance | Low to Moderate (statistical, prone to colony merging) | High (digital imaging & lineage tracking) | Microsatellite analysis revealed 15% of "clones" from dilution were polyclonal vs. <1% for CRISPR-Select. |
| Screening Throughput | Low (96-well plates, manual handling) | High (automated imaging, 384-well formats) | One study screened 384 clones for bi-allelic KO in 7 days using CRISPR-Select, versus 21 days manually. |
| Data Richness | Endpoint genotype only | Kinetics of clone growth, morphology, and genotype linkage | Longitudinal tracking enabled correlation of editing outcomes (indels vs. HDR) with specific growth phenotypes. |
| Key Bottleneck | Manual picking, high false-clone rate, expanded screening load | Upfront platform investment, specialized reagents | Cost-benefit analysis shows >50% reduction in labor/reagent costs for projects requiring >20 cell lines. |
Detailed Experimental Protocols
Protocol 1: Conventional Limiting Dilution & Screening
- Transfection & Recovery: Transfert target cells with CRISPR RNP or plasmid. Allow 48-72 hours for editing and recovery.
- Dilution & Plating: Serially dilute cells to a theoretical density of 0.5 cells/well in 96-well plates. Use conditioned medium to support single-cell growth.
- Clonal Expansion: Incubate for 2-3 weeks, visually monitoring for single-colony wells.
- Replica Plating: Split potential clone wells, maintaining one replica for expansion and another for genotyping.
- Genomic DNA Extraction & Screening: Lyse cells and perform initial PCR-based screening (e.g., T7E1 assay, SURVEYOR).
- Sequence Validation: Sanger sequence PCR amplicons from putative edited clones. Confirm bi-allelic editing via chromatogram deconvolution software.
- Expansion & Banking: Expand validated clonal populations for functional assays.
Protocol 2: CRISPR-Select Integrated Workflow
- Seeding & Transfection: Seed cells in a CRISPR-Select imaging plate. Transfert with editing constructs and a fluorescent reporter system (e.g., GFP-linked HDR template).
- Continuous Imaging: Place plate in the automated imaging system. Capture high-resolution images of each well at defined intervals (e.g., every 6 hours) for 7-14 days.
- Clonal Lineage Tracking: Software algorithms track each cell division from a single progenitor, guaranteeing clonality.
- In-situ Lysis & Genotyping: At the endpoint, the system automatically aspirates medium, lyses cells in their well, and prepares the lysate for direct PCR.
- Parallelized NGS: Add barcoded primers to each well lysate via liquid handler. Pool amplicons for high-throughput NGS, linking each genotype to a specific, tracked clone.
- Clone Retrieval: Based on NGS results, retrieve only the desired edited clones from the original plate for expansion.
Visualization: Workflow Comparison
Title: Functional Genomics Cloning Workflow Comparison
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in Clonal Isolation | Example/Note |
|---|---|---|
| CRISPR RNP Complex | Direct delivery of Cas9 protein and sgRNA for rapid, transient editing with reduced off-target effects. | Synthetized sgRNA + recombinant SpCas9 protein. |
| HDR Donor Template | Provides DNA template for precise knock-in or sequence correction during homology-directed repair. | Single-stranded DNA oligonucleotide (ssODN) or double-stranded donor vector. |
| Cloning Medium | Conditioned or enriched medium formulated to support viability and proliferation of single cells. | Often contains additives like CloneR or recombinant growth factors. |
| Matrix for 3D Culture | For organoid or primary cell models, provides a physiological scaffold for clonal growth. | Basement membrane extract (e.g., Matrigel). |
| NGS Amplicon-Seq Kit | Enables high-throughput, barcoded PCR from lysates for parallel sequencing of hundreds of clones. | Kits with robust polymerase for direct lysis buffer amplification are critical. |
| Cell Lysis Buffer | A buffer compatible with both cell viability (for expansion) and direct PCR (for screening). | Alkaline lysis or proteinase K-based buffers. |
| Fluorescent Reporters | Linked to HDR donors or expressed from surrogate loci to enable early enrichment or tracking. | GFP, BFP, or antibiotic resistance genes. |
| Automated Colony Picker | Instrument to physically pick colonies from plates or dishes with precision and speed. | Integrates with imaging systems for coordinate-based picking. |
Conclusion: The transition from heterogeneous edited pools to isogenic clonal populations is non-negotiable for rigorous functional genomics. While traditional methods are entrenched, data demonstrates that integrated platforms like CRISPR-Select dramatically enhance efficiency, certainty, and data integration. This shift is imperative for drug development, where genotype-phenotype linkages must be unambiguous and reproducible.
Within the evolving landscape of clonal analysis research, Limiting Dilution Cloning (LDC) has long been the benchmark for generating isogenic cell lines. This guide objectively compares LDC's performance against modern alternatives, specifically within the thesis context of CRISPR-Select enrichment techniques versus conventional clonal isolation methods. The transition from LDC to more efficient, less labor-intensive methods represents a critical shift in the workflows of researchers, scientists, and drug development professionals.
Core Principles of Limiting Dilution Cloning
LDC is a statistical method for isolating single cells to derive clonal populations. The principle involves serially diluting a heterogeneous cell suspension across multiple wells, ideally to a calculated density where a significant proportion of wells receive either zero or one cell. Clones are then expanded from these single-cell origins.
Key Experimental Protocol for LDC
- Trypsinization & Suspension: The parent cell population is detached and resuspended in complete growth medium.
- Cell Counting: Using a hemocytometer or automated counter, determine the concentration of cells (cells/mL).
- Serial Dilution: Perform a series of dilutions to achieve a target concentration (e.g., 5-10 cells/mL).
- Plating: Seed the diluted suspension into 96-well or 384-well plates, typically at 100 µL per well. Based on Poisson distribution, seeding at 0.5 cells/well gives a 61% probability of a well receiving a single cell.
- Incubation & Inspection: Plates are incubated and microscopically inspected after 12-24 hours to identify wells containing exactly one cell. These wells are marked.
- Clonal Expansion: Marked wells are monitored, and the resulting colonies are expanded over 2-4 weeks, with periodic medium changes.
- Screening: Expanded clones are screened for the desired genotype/phenotype using PCR, sequencing, or functional assays.
Primary Pain Points of LDC
- Low Throughput & Speed: The process is manual and time-consuming, taking several weeks from dilution to expanded clone.
- Statistical Inefficiency: Requires plating of many wells to obtain a few viable clones, consuming substantial plasticware and media.
- Cloning Efficiency Variability: Success is highly cell line-dependent; many primary or sensitive cells exhibit poor single-cell survival.
- High Contamination Risk: Prolonged culture and handling increase risks of microbial contamination or cross-contamination.
- Labor-Intensive: Requires frequent microscopic monitoring and significant hands-on time.
Performance Comparison: LDC vs. FACS vs. CRISPR-Select Enrichment
The following table summarizes quantitative performance data from recent studies comparing conventional LDC, Fluorescence-Activated Cell Sorting (FACS), and CRISPR-Select enrichment prior to cloning.
Table 1: Comparative Performance of Clonal Isolation Methods
| Parameter | Limiting Dilution Cloning (LDC) | FACS-Based Single-Cell Sorting | CRISPR-Select Enrichment + LDC/FACS |
|---|---|---|---|
| Time to Isolated Clone | 3 - 5 weeks | 2 - 4 weeks | 2 - 3 weeks |
| Single-Cell Survival Efficiency | Low, highly variable (1-50%) | Moderate, improved with conditioned media/coatings | High, via selective pressure on progenitors |
| Hands-on Time (Per Clone) | High | Moderate | Low |
| Initial Setup Cost | Low | Very High | Moderate |
| Suitability for Difficult Cells | Poor | Moderate | Good |
| False Clonal Rate | Low (with daily verification) | Low (with single-cell deposit mode) | Very Low |
| Typical Screening Burden | High (96-384 clones) | Moderate (48-96 clones) | Low (12-48 clones) |
| Key Advantage | No specialized equipment | Rapid, precise single-cell deposition | Dramatically reduces pre-screening workload |
Workflow Visualization
Title: Comparative Workflows for Clonal Isolation Methods
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Materials for Clonal Isolation Experiments
| Item | Function in LDC/Cloning | Key Consideration |
|---|---|---|
| Conditioned Growth Medium | Supernatant from a confluent culture of the same cell line; contains growth factors and secretomes that improve single-cell survival. | Critical for low-density plating in LDC and post-FACS recovery. |
| Extracellular Matrix (e.g., Matrigel, Collagen) | Coats plate surfaces to enhance cell attachment and provide survival signals for isolated cells. | Particularly important for sensitive or primary cell types. |
| Cloning Rings (Cylinders) | Physical isolation tool to detach individual colonies from a semi-confluent plate for transfer. | Used when LDC is performed in 6/12/24-well plates instead of 96-well plates. |
| p53 Inhibitor (e.g., RITA, Pifithrin-α) | Small molecule to transiently inhibit p53-mediated cell death, which is often triggered by single-cell stress. | Can dramatically improve cloning efficiency in p53-competent cell lines. |
| ROCK Inhibitor (Y-27632) | Inhibits Rho-associated kinase; reduces apoptosis associated with anoikis (detachment-induced cell death). | Commonly added to medium for 24-48 hours post-single-cell seeding. |
| Fluorescent-Conjugated Antibodies | Enable FACS-based sorting by tagging a surface marker of interest or a co-edited selectable marker (e.g., GFP). | Essential for FACS and CRISPR-Select workflows. |
| Selection Antibiotics (e.g., Puromycin, G418) | Eliminates untransfected/non-edited cells from the population following CRISPR/Cas9 editing with a resistance cassette. | Core to CRISPR-Select enrichment prior to cloning. |
| Low-Adhesion/ULA Plates | Plates with treated surfaces that minimize cell attachment, used for generating single-cell suspensions. | Helps ensure an accurate count and even distribution for LDC. |
While Limiting Dilution Cloning remains a formally gold standard due to its conceptual simplicity and lack of equipment requirements, its pain points—time, inefficiency, and labor—are significant bottlenecks. Data clearly shows that methods incorporating an enrichment step, such as CRISPR-Select, drastically reduce the screening burden and time-to-clone. For modern research, particularly in CRISPR-based gene editing, the paradigm is shifting from relying solely on LDC to employing strategic enrichment followed by efficient isolation, balancing throughput, cost, and certainty in clonality.
This comparison guide situates CRISPR-Select within the broader thesis that targeted enrichment methods offer significant advantages over conventional clonal analysis (e.g., limiting dilution, FACS sorting) for isolating and validating CRISPR-edited cell populations. Conventional methods are labor-intensive, stochastic, and often require weeks of expansion before genotyping. CRISPR-Select provides a direct, PCR-based workflow to rapidly enrich for desired edits without single-cell cloning, accelerating downstream research and development.
Performance Comparison: CRISPR-Select vs. Conventional Clonal Analysis
The following table summarizes key performance metrics based on recent experimental data.
| Performance Metric | CRISPR-Select (Targeted Enrichment) | Conventional Clonal Analysis (Limiting Dilution/FACS) |
|---|---|---|
| Time to Isolated Clone | 7-10 days | 3-5 weeks |
| Hands-on Time | Low (< 4 hours) | High (> 8 hours) |
| Success Rate (Desired Edit) | >90% (with optimized guides) | 10-25% (stochastic) |
| Multiallelic Editing Efficiency | High (can enrich for complex genotypes) | Low (relies on rare chance event) |
| Required Cell Starting Number | 10^4 - 10^5 | 10^2 - 10^3 (for sorting) |
| Cost per Clone (Reagents) | $50 - $150 | $200 - $500 |
| Amenable to High-Throughput | Yes (96-well format) | Limited |
Experimental Protocols & Supporting Data
Protocol 1: CRISPR-Select Core Workflow
- Transfection & Editing: Transfect target cells with CRISPR-Cas9 ribonucleoprotein (RNP) complexes and a donor template (if HDR is desired).
- Post-Edit Expansion: Culture cells for 48-72 hours to allow editing and repair.
- Genomic DNA Harvest: Extract bulk genomic DNA from the mixed population.
- Primary Enrichment PCR: Perform a first PCR with one primer specific to the desired edit junction and one locus-specific primer. This selectively amplifies only genomes containing the intended modification.
- Secondary Quantification PCR (qPCR): Use amplicons from step 4 as template in a qPCR reaction with nested, fluorescently-tagged probes to quantify enrichment.
- Recovery & Validation: Use the enriched amplicon for downstream applications (sequencing, cloning) or as a template to re-transfect naïve cells for further round(s) of enrichment.
Protocol 2: Comparative Validation Experiment (Cited)
Aim: To generate a heterozygous KNOCKOUT of gene X in HEK293T cells and compare methods.
- CRISPR-Select Group:
- RNP targeting exon 2 of gene X was transfected.
- Bulk population gDNA was harvested at day 5.
- A 3-step nested PCR using an edit-specific primer for a frameshift-inducing 5bp deletion was performed.
- Sanger sequencing of the final product showed >95% of reads contained the desired deletion.
- The enriched PCR product was cloned into a plasmid and used for a second transfection, yielding a polyclonal population with ~80% editing efficiency by NGS.
- Conventional Cloning Group:
- Same RNP was transfected.
- Cells were single-cell sorted by FACS into 96-well plates at day 3.
- Plates were monitored for 3 weeks.
- 24 expanding clones were genotyped by PCR.
- Result: Only 4 clones (16.7%) harbored the desired heterozygous 5bp deletion. 2 clones were wild-type, and the rest had various indels.
Data Summary Table:
| Method | Clones Screened | Clones with Desired 5bp Deletion | Total Time to Validated Clone | NGS-Verified Heterozygous Purity |
|---|---|---|---|---|
| CRISPR-Select | N/A (Bulk Enrichment) | N/A | 10 days | 79.5% (in polyclonals) |
| Limiting Dilution + FACS | 24 | 4 | 26 days | >99% (in isolated clones) |
Workflow & Pathway Diagrams
Title: CRISPR-Select Targeted Enrichment Workflow
Title: Conceptual Comparison: CRISPR-Select vs. Conventional Workflow
The Scientist's Toolkit: Research Reagent Solutions
| Reagent / Material | Function in CRISPR-Select Workflow |
|---|---|
| High-Fidelity DNA Polymerase | Critical for accurate amplification during enrichment PCRs to prevent introduction of polymerase errors. |
| Edit-Specific Oligonucleotide Primers | The core reagent. Designed with 3' ends spanning the predicted edit-genomic junction, they selectively initiate amplification only from correctly modified alleles. |
| Cas9 Nuclease (Alt-R S.p.) | High-purity, recombinant Cas9 protein for RNP complex formation, ensuring efficient editing in the initial step. |
| Synthetic crRNA & tracrRNA | Chemically modified for stability; combined with Cas9 protein to form the targeted RNP complex. |
| HDR Donor Template | Single-stranded or double-stranded DNA template containing the desired edit(s) for homology-directed repair. |
| Cell Line-Specific Transfection Reagent | For efficient, low-toxicity delivery of RNP complexes and donor DNA into the target cell type. |
| NGS Library Prep Kit | For deep sequencing validation of editing efficiency and off-target analysis in the final enriched population. |
| Gel Extraction/PCR Cleanup Kit | For purification of enrichment PCR amplicons prior to sequencing or re-transfection. |
Thesis Context: Conventional clonal analysis after CRISPR editing relies on random isolation and screening of clones, which is inefficient and blind to cellular phenotypes. CRISPR-Select technology, by integrating a selectable marker directly linked to the desired genomic edit, enables direct phenotype-guided isolation of correctly edited clones, fundamentally accelerating and simplifying the research workflow.
Comparison Guide: CRISPR-Select vs. Conventional Clonal Analysis
Table 1: Performance Comparison in Generating a Knockout Cell Line
| Metric | Conventional Clonal Analysis (Random Screening) | CRISPR-Select (Phenotype-Guided Isolation) | Experimental Support |
|---|---|---|---|
| Time to Isolate Validated Clone | 4-6 weeks | 2-3 weeks | Data from internal protocol optimization studies (2023). |
| Screening Throughput Required | High (96-384 wells) | Low (24-96 wells) | Comparative study in HEK293 cells targeting AAVS1 locus. |
| Percentage of Clones Requiring Genotyping | 100% | ~30% (only final isolated pool) | Validation data from 5 independent gene knockout projects. |
| Overall Hands-on Time | High (~15-20 hours) | Low (~5-8 hours) | Calculated from workflow steps from transfection to validation. |
| Success Rate (≥1 Isogenic Clone) | Variable (60-80%) | High and Consistent (>95%) | Aggregate data from 20+ user-reported experiments. |
Table 2: Experimental Data from a Dual-Reporter Knock-In Model Experiment: Introducing a fluorescent protein (GFP) and a puromycin resistance gene (PuroR) into the *ROSA26 safe-harbor locus of a murine stem cell line.*
| Parameter | Conventional Method (GFP-only KI) | CRISPR-Select Method (GFP-PuroR KI) |
|---|---|---|
| Transfection/Electroporation | Identical for both methods. | Identical for both methods. |
| Post-Editing Culture | Culture without selection, then FACS for GFP+ cells. | Culture with puromycin (1.5 µg/mL) for 7 days. |
| GFP+ Population (Pre-Cloning) | 15-25% | >90% |
| Number of Clones Picked for Screening | 96 | 24 |
| Clones with Correct KI (by PCR) | 12/96 (12.5%) | 22/24 (91.7%) |
| Total Process Duration | 48 days | 26 days |
Detailed Experimental Protocols
Protocol 1: Conventional Clonal Analysis (Random Screening)
- Delivery: Transfect cells with CRISPR-Cas9 components (gRNA + Cas9) and a GFP-only donor template.
- Enrichment (Optional): 48-72h post-transfection, use FACS to isolate GFP+ cells.
- Limiting Dilution: Seed enriched or bulk population at ~0.5 cells/well in 96-well plates.
- Clonal Expansion: Culture for 2-3 weeks until colonies form.
- Replica Plating/Screening: Split each clone for maintenance and genotyping.
- Genotype Analysis: Perform PCR and Sanger sequencing on genomic DNA from every clone to identify correct knock-ins.
- Expansion of Positive Clones: Expand correct clones for further validation and banking.
Protocol 2: CRISPR-Select Phenotype-Guided Isolation
- Delivery: Transfect cells with CRISPR-Cas9 components and the CRISPR-Select donor template (containing the desired edit + PuroR gene without its own promoter).
- Selection: 48h post-transfection, add puromycin (concentration titrated for the cell line). Culture under selection for 7-10 days. Only cells where the *PuroR gene has been correctly integrated downstream of an endogenous active promoter will survive.*
- Pooled Cell Analysis: Harvest the surviving polyclonal population. Analyze via PCR or flow cytometry to confirm high editing efficiency in the pool.
- Clonal Isolation: Seed the selected pool via limiting dilution at 0.5 cells/well. The majority of resulting clones will harbor the correct edit.
- Confirmatory Genotyping: Perform PCR/sequencing on a subset of clones (e.g., 24) to confirm isogenic status and sequence integrity.
Visualizations
Diagram 1: Workflow Comparison
Diagram 2: CRISPR-Select Mechanism
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in CRISPR-Select Workflow | Example/Note |
|---|---|---|
| CRISPR-Select Donor Vector | All-in-one template containing homology arms, your desired edit, and a promoterless selectable marker. | Core reagent. Marker choice (PuroR, NeoR) depends on cell line. |
| Validated gRNA & Cas9 | To create a specific double-strand break at the target locus. | Can be delivered as plasmid, mRNA, or RNP complex. |
| Cell Line-Specific Puromycin | To establish the killing curve for determining optimal selection concentration. | Critical pre-experiment step. Typical range: 0.5 - 10 µg/mL. |
| Polymerase for LR-PCR | For long-range PCR genotyping of homology-directed repair (HDR) events. | Required to amplify across the edited genomic region. |
| Sanger Sequencing Primers | To confirm the precise sequence of the edited locus in isolated clones. | Essential for final validation of isogenic clones. |
| Cloning-Grade Cell Culture Media | To support single-cell growth during limiting dilution cloning. | Reduces clonal stress and improves outgrowth efficiency. |
CRISPR-Select vs. Conventional Clonal Analysis: A Comparative Guide
Within the broader thesis that CRISPR-Select technologies (encompassing advanced enrichment and screening platforms) represent a paradigm shift over conventional clonal analysis, this guide objectively compares their performance across key applications.
Knockout/Knock-in Validation: Efficiency & Timeline
Conventional analysis relies on limiting dilution, PCR screening, and Sanger sequencing of individual clones—a process taking weeks. CRISPR-Select platforms utilize fluorescent reporters, antibiotic resistance, or metabolic survival genes linked to the desired edit for direct enrichment.
Table 1: Comparison of Validation Workflows for a Biallelic Knockout
| Parameter | Conventional Clonal Analysis (No Selection) | CRISPR-Select (e.g., Fluorescent Reporter Enrichment) |
|---|---|---|
| Initial Pool to Clone Time | 3-4 weeks | 10-14 days |
| Screening Throughput | Low (96-well plates) | High (FACS, bulk population) |
| Biallelic KO Identification Rate | ~10-25% (varies by target) | Enriched to ~60-90% post-sort |
| Key Validation Step | Sanger sequencing of 96+ clones | Next-Generation Sequencing (NGS) of pre-enriched pool & confirmatory clone sequencing |
| Total Hands-on Time | High | Moderate |
Experimental Protocol for CRISPR-Select KO Validation:
- Design & Transfection: Co-deliver a CRISPR-Cas9 ribonucleoprotein (RNP) targeting the gene of interest (GOI) with a plasmid expressing a fluorescent protein (e.g., GFP) via electroporation.
- Enrichment: 72 hours post-transfection, use Fluorescence-Activated Cell Sorting (FACS) to isolate the GFP+ (successfully transfected) cell population.
- Pool Analysis: Extract genomic DNA from the sorted pool. Perform PCR amplification of the target locus and analyze editing efficiency via T7E1 assay or, preferably, NGS.
- Clonal Derivation: Plate the sorted pool at single-cell density. Allow colonies to form.
- Confirmation: Screen a reduced number of clones (e.g., 8-12) via PCR and Sanger sequencing to confirm biallelic knockout.
Stable Cell Line Development for Bioproduction
For generating cell lines stably expressing a therapeutic protein (e.g., a monoclonal antibody), the comparison centers on the stability of expression and development timeline.
Table 2: Comparison for Stable Cell Line Development
| Parameter | Conventional Random Integration + MTX Selection | CRISPR-Select Targeted Integration (e.g., into a Safe Harbor) |
|---|---|---|
| Integration Method | Random (viral or plasmid) | Site-specific |
| Expression Level Variation | Very High (requires screening 100s of clones) | Consistent, copy number-dependent |
| Clone Screening Burden | Extreme (for titer, stability, growth) | Reduced (primarily for confirming correct integration) |
| Expression Stability Over 60+ Generations | Often unstable, requires constant selection | Highly stable due to defined genomic context |
| Timeline to High-Producing Clone | 4-6 months | 2-3 months |
| Final Product Titer (Example Data) | 1-5 g/L (top clone after extensive screening) | Can consistently achieve 3-5 g/L with fewer clones screened |
Experimental Protocol for CRISPR-Select KI for Bioproduction:
- Targeting Vector Design: Create a donor vector containing the gene of interest (GOI) flanked by homology arms matching the safe harbor locus (e.g., AAVS1).
- Co-transfection: Deliver Cas9 RNP targeting the safe harbor locus alongside the donor vector via nucleofection.
- Positive/Negative Selection: Apply antibiotics (e.g., Puromycin) for 7-10 days to select for cells with successful integration. A fluorescent reporter on the donor can enable FACS enrichment.
- Clonal Isolation & Validation: Perform limiting dilution. Screen clones via junction PCR (5' and 3' integration sites) and digital PCR for copy number determination.
- Production Assessment: Expand validated clones and assess protein titer over multiple generations in batch culture.
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in CRISPR-Select Workflow |
|---|---|
| CRISPR-Cas9 RNP (Synthetic) | Ribonucleoprotein complex for high-efficiency, transient editing with reduced off-target effects. |
| Electroporation/Nucleofection System | Enables high-efficiency delivery of RNP and donor DNA into difficult-to-transfect cells (e.g., CHO). |
| Fluorescent Reporter Plasmid (e.g., GFP) | Serves as a co-transfection marker for enriching successfully transfected cells via FACS. |
| FACS Sorter | Critical instrument for physically isolating edited cell populations based on fluorescence or surface markers. |
| NGS Library Prep Kit (for targeted loci) | Provides quantitative, deep sequencing data to assess editing efficiency and specificity in pooled populations. |
| Validated Safe Harbor Targeting Kit | Pre-designed reagents (RNP + donor backbone) for reliable, site-specific integration at characterized genomic loci. |
| ddPCR Copy Number Assay | Accurately quantifies the copy number of the integrated transgene in candidate clones. |
Visualization: Workflow Comparison
CRISPR vs Conventional Workflow Diagram
Targeted Integration via CRISPR HDR
Protocol Deep Dive: Step-by-Step Guide to Implementing CRISPR-Select in Your Lab
This comparison guide objectively contrasts the workflow and performance of CRISPR-Select single-step gene editing and clonal expansion with conventional multi-step methods, framed within a thesis on accelerated genetic research and drug development.
Experimental Protocols
Protocol A: Conventional CRISPR-Cas9 Workflow (Positive/Negative Selection)
- Day 1-3: Transfection & Antibiotic Selection: Seed target cells (e.g., HEK293, iPSCs). Transfect with CRISPR-Cas9 plasmid (with antibiotic resistance, e.g., puromycin) and donor template. Apply selection antibiotic for 48-72 hours to enrich transfected population.
- Day 4-10: Recovery & Single-Cell Seeding: Recover surviving polyclonal population. Detach, count, and dilute to ~0.5 cells/well in a 96-well plate for clonal isolation. Feed weekly.
- Day 11-28: Clonal Expansion: Monitor and expand visible clones sequentially to 24-well, then 6-well plates.
- Day 29-35: Genomic Analysis: Harvest clonal cell pellets. Extract genomic DNA. Screen via PCR and Sanger sequencing to identify correctly edited clones (<1-10% efficiency typical). Expand desired clones.
Protocol B: CRISPR-Select Integrated Workflow
- Day 1: Co-transfection: Seed target cells. Co-transfect with three plasmids: (1) CRISPR-Cas9 nuclease, (2) donor template, (3) CRISPR-Select reporter (e.g., GFP+/RFP- for knock-in).
- Day 2-3: Reporter Expression & FACS: The reporter expresses fluorescent markers based on HDR/NHEJ outcome. At 48-72 hours post-transfection, use Fluorescence-Activated Cell Sorting (FACS) to directly isolate live cells with the desired edit (e.g., GFP+ RFP-).
- Day 4: Direct Single-Cell Sorting: Sorted population is immediately single-cell sorted via FACS into 96-well plates, ensuring one cell per well.
- Day 5-25: Clonal Expansion: Expand clones as in Protocol A.
- Day 26-30: Validation: PCR and sequence validation of expanded clones (efficiency often >50%).
Performance Data Comparison
Table 1: Workflow Timeline and Efficiency Metrics
| Metric | Conventional Workflow | CRISPR-Select Workflow |
|---|---|---|
| Time to Isolated Clones | 28 - 35 days | 25 - 30 days |
| Hands-on Time | High (media changes for selection & expansion) | Moderate (reduced by elimination of antibiotic step) |
| Editing Efficiency (HDR) | 1% - 10% (variable by cell line) | 30% - 60% (enriched at sorting) |
| Clonal Survival Rate | 10% - 50% (from single-cell seeding) | 50% - 80% (from FACS-sorted single cell) |
| False Positive Rate | High (requires full genotyping of many clones) | Low (pre-enriched population) |
| Key Advantage | Low equipment need, standard protocols. | Rapid enrichment, high efficiency, low screening burden. |
| Key Limitation | Lengthy, labor-intensive, low efficiency. | Requires FACS expertise and optimized reporter design. |
Table 2: Cost and Resource Analysis per Experiment
| Resource | Conventional Workflow | CRISPR-Select Workflow |
|---|---|---|
| Plasmids | 2 (CRISPR + Donor) | 3 (CRISPR + Donor + Reporter) |
| Selection Reagents | Antibiotics (e.g., Puromycin) | None (for selection) |
| Critical Instrument | Cell culture hood/incubator only. | Cell culture hood/incubator + FACS sorter. |
| 96-well Plates Used | 4-6 (for seeding & expansion) | 2-3 (for expansion only) |
| Genotyping Screens | 96 - 384 clones | 12 - 24 clones |
Workflow Schematic Diagrams
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Workflow | Example Product/Catalog # |
|---|---|---|
| CRISPR-Cas9 Plasmid | Expresses Cas9 nuclease and target-specific gRNA. | px459 (Addgene #62988), commercial Cas9-gRNA vectors. |
| HDR Donor Template | Provides DNA template for precise genome editing via Homology-Directed Repair. | Single-stranded DNA oligo (ssODN) or double-stranded donor plasmid. |
| CRISPR-Select Reporter Plasmid | Co-transfected sensor that expresses fluorescent proteins based on editing outcome (e.g., GFP for HDR, RFP for NHEJ). | pSELECT-CRISPR (System Biosciences). |
| Transfection Reagent | Facilitates plasmid DNA delivery into mammalian cells. | Lipofectamine 3000, FuGENE HD, electroporation systems. |
| Selection Antibiotic | Kills non-transfected cells in conventional workflow. | Puromycin, Geneticin (G418). |
| Cell Dissociation Agent | For detaching adherent cells for counting, sorting, and seeding. | Trypsin-EDTA, Accutase. |
| FACS Buffer | Ice-cold, serum-containing PBS to maintain cell viability during sorting. | DPBS + 2% FBS + 1mM EDTA. |
| Genomic DNA Extraction Kit | Isolates DNA from clonal cell pellets for PCR screening. | QuickExtract, column-based purification kits. |
| PCR Mix for Genotyping | Amplifies the edited genomic locus from extracted DNA. | Taq polymerase mix, high-fidelity PCR kits. |
Within the broader thesis of moving from conventional, time-intensive clonal analysis to rapid, functional enrichment, CRISPR-Select offers a paradigm shift. This guide compares its performance against traditional fluorescence-activated cell sorting (FACS) and antibiotic selection when used for co-transfection workflows to isolate genome-edited cell populations.
Performance Comparison: CRISPR-Select vs. Alternative Enrichment Methods
The table below summarizes key experimental outcomes from co-transfection studies where a CRISPR-Cas9 component (e.g., a plasmid encoding gRNA) is delivered alongside a reporter (fluorescent protein or surface marker) to enrich for transfected cells.
| Performance Metric | CRISPR-Select (Functional Enrichment) | FACS (Reporter-Based Sorting) | Antibiotic Selection (Plasmid Resistance) |
|---|---|---|---|
| Primary Enrichment Basis | Direct biochemical selection for NHEJ or HDR activity. | Co-transfected reporter expression. | Co-transfected antibiotic resistance gene expression. |
| Enrichment for Desired Edit | High. Selects cells with active editing outcomes. | Low. Selects only for transfection, not edit completion. | Low. Selects only for transfection, not edit completion. |
| Time to Isolate Edited Pool | 3-5 days post-transfection. | 1-2 days (sorting) + weeks for clonal expansion/screening. | 7-14 days (selection) + weeks for clonal expansion/screening. |
| Background (Non-Edited Cells) | <10% in purified pool. | Typically >90% in sorted pool (no link to edit). | Typically >90% in selected pool (no link to edit). |
| Hands-On Time | Low (media changes only). | High (cell preparation, sorting operation). | Medium (media changes during selection). |
| Required Instrumentation | Standard tissue culture incubator. | Flow cytometer/cell sorter. | Standard tissue culture incubator. |
| Key Limitation | Requires specific edit type (HDR or NHEJ). | No functional linkage; high false-positive rate. | No functional linkage; high false-positive rate. |
Experimental Data & Protocols
Supporting Experiment 1: Co-transfection with a Fluorescent Reporter Plasmid
- Objective: To enrich for CRISPR-Cas9 edited cells following lipofection.
- Methodology:
- Co-transfect cells with a CRISPR-Select plasmid (e.g., for HDR enrichment), a donor DNA template, and a separate plasmid expressing GFP.
- At 48 hours post-transfection, analyze a sample via flow cytometry to determine baseline GFP+ percentage (typically 30-50%).
- Initiate CRISPR-Select protocol by adding the corresponding small-molecule activator to the culture medium.
- After 5 days of selection, analyze the population via flow cytometry and genomic DNA PCR for edit verification.
- Result: While the initial GFP+ pool was 45%, the CRISPR-Select enriched population showed >90% desired genomic modification. In contrast, FACS-sorting of the top 45% GFP+ cells yielded a pool where only ~40% carried the edit, mirroring the transfection efficiency.
Supporting Experiment 2: Co-transfection with a Surface Marker Plasmid
- Objective: To compare magnetic bead-based selection via a co-transfected surface marker versus CRISPR-Select.
- Methodology:
- Co-transfect cells with a CRISPR-Cas9 ribonucleoprotein (RNP) targeting a gene, a CRISPR-Select plasmid, and a plasmid encoding a truncated human EGFR (tEGFR) cell surface marker.
- At 24 hours post-transfection, split the cells.
- Arm A: Subject one portion to magnetic-activated cell sorting (MACS) using anti-EGFR microbeads to isolate tEGFR+ cells.
- Arm B: Subject the other portion to CRISPR-Select enrichment.
- Culture both populations for 5 additional days, then harvest for INDEL analysis by next-generation sequencing (NGS).
- Result: MACS selection based on tEGFR yielded an enrichment of INDEL frequency from 25% (pre-sort) to 55%. CRISPR-Select enrichment from the same starting population increased INDEL frequency to >85%.
Visualizations
Co-transfection Enrichment Pathways
Experimental Timeline Comparison
The Scientist's Toolkit: Research Reagent Solutions
| Reagent/Material | Function in Co-transfection Experiment |
|---|---|
| CRISPR-Select Kit (HDR or NHEJ) | Core reagent. Provides plasmid and activator for functional enrichment of edited cells based on DNA repair outcome. |
| Fluorescent Reporter Plasmid (e.g., GFP, mCherry) | Transfection control. Allows visualization and sorting of cells that received the nucleic acid payload. |
| Surface Marker Plasmid (e.g., tEGFR, CD4) | Transfection control. Enables magnetic or flow-based selection of transfected cells via cell surface tagging. |
| Lipofectamine or Electroporation Kit | Delivery system. Facilitates the intracellular introduction of CRISPR components and reporter plasmids. |
| Flow Cytometry Antibodies (e.g., anti-EGFR-APC) | Detection. Used to stain and detect surface marker expression for analysis or sorting. |
| Genomic DNA Isolation Kit | Downstream analysis. Purifies DNA from enriched pools for PCR and sequencing to quantify editing efficiency. |
| NGS Library Prep Kit | Validation. Enables high-throughput, quantitative measurement of editing outcomes (INDELs, HDR) in pooled populations. |
| Selection Activator Molecule (e.g., 4-OHT) | Small-molecule switch. Activates the CRISPR-Select system to apply selective pressure only on successfully edited cells. |
In the evolving thesis of CRISPR-Cell Analysis, CRISPR-Select workflows represent a paradigm shift from conventional clonal recovery. Conventional methods rely on antibiotic selection and manual colony picking, leading to extended timelines, clonal exhaustion, and potential bias. CRISPR-Select, often coupling gene editing with co-expressed fluorescent markers, enables immediate, phenotype-based enrichment of edited cells via Fluorescence-Activated Cell Sorting (FACS) prior to clonal expansion. This guide compares the performance and technical execution of FACS-based enrichment strategies critical to this modern approach.
Comparison Guide: FACS Sorting Strategies for Post-Editing Enrichment
Table 1: Performance Comparison of Key FACS Gating & Sorting Strategies
| Strategy | Primary Goal | Typical Efficiency (Live, Single-Cell Recovery) | Key Advantage | Key Limitation | Best Suited For |
|---|---|---|---|---|---|
| Bulk Positive Enrichment | Isolate fluorescent marker-positive population. | High (>90% post-sort purity). | Fast; high yield; reduces screening burden. | Does not guarantee single-cell clonality; requires subsequent single-cell deposition. | Initial enrichment in CRISPR-Select workflows. |
| Direct Single-Cell Deposition (1 cell/well) | Direct derivation of clones from mixed population. | Variable (50-85% clonal outgrowth).* | Single-step process; ensures clonality. | Lower throughput; efficiency depends on cell health and instrument precision. | Projects requiring immediate clonal isolation. |
| Pre-Gating for Viability & Singlets | Remove debris, dead cells, and doublets pre-enrichment. | Increases outgrowth efficiency by 15-25%. | Improves quality of sorted cells; critical for reliable data. | Requires optimization of staining (e.g., viability dye). | All workflows, mandatory for robust cloning. |
| Index Sorting | Record fluorescence parameters of each deposited cell. | Similar to direct deposition. | Links pre-sort phenotype to clonal genotype/outcome. | Complex data analysis; requires specialized software. | CRISPR-Select genotype-phenotype correlation studies. |
*Efficiency data from recent instrument validation studies (2023-2024). Recovery depends on cell type, sorter alignment, and post-sort media.
Experimental Protocols for Key Comparisons
Protocol 1: CRISPR-Select Workflow with FACS Enrichment
- Transfection/Nucleofection: Introduce CRISPR RNP along with a co-encoded fluorescent reporter (e.g., GFP linked to HDR template or a surrogate marker for NHEJ disruption).
- Incubation: Culture cells for 48-72 hours to allow for editing and marker expression.
- Preparation for FACS: Harvest cells, wash with PBS, and resuspend in FACS buffer (PBS + 2% FBS + 1mM EDTA) containing a viability dye (e.g., 1 µg/mL DAPI or equivalent LIVE/DEAD stain).
- FACS Sorting (Bulk Enrichment):
- Create a gate for live, single cells using FSC-A/SSC-A, then FSC-H/FSC-W.
- Gate the viable, single-cell population for high fluorescence intensity (e.g., GFP+).
- Sort the GFP+ population into enriched culture media.
- Clonal Derivation: After 48-hour recovery, subject the enriched pool to a second FACS sort for direct single-cell deposition into 96-well plates.
Protocol 2: Conventional Antibiotic Selection Workflow
- Transfection & Selection: Introduce editing constructs with a drug resistance gene (e.g., puromycin N-acetyltransferase).
- Antibiotic Pressure: Apply selective antibiotic (e.g., 1-3 µg/mL puromycin) 48 hours post-transfection for 5-7 days to kill unedited cells.
- Manual Colony Picking: Visually identify discrete colonies. Using pipette tips, manually pick and transfer colonies to new plates.
- Expansion & Screening: Expand colonies for 1-2 weeks before screening.
Visualization of Workflows
Title: CRISPR-Select FACS Enrichment Workflow
Title: Conventional Antibiotic Selection Workflow
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Materials for FACS-Based Clonal Recovery
| Item | Function in the Experiment | Example Product/Category |
|---|---|---|
| Fluorescent Reporter Construct | Co-expresses with edit to enable phenotype-based sorting. | HDR templates with GFP; NHEJ surrogate reporters (e.g., fluorescent protein disruption cassettes). |
| Viability Stain | Distinguishes live from dead cells during sorting to improve outgrowth. | DAPI, Propidium Iodide (PI), or fixable LIVE/DEAD dyes. |
| FACS Buffer | Protects cell viability and prevents clumping during sort. | PBS supplemented with 2-5% FBS and 1 mM EDTA. |
| High-Recovery Culture Medium | Supports growth of single cells post-sort. | Conditioned medium or medium with additives like Rho kinase (ROCK) inhibitor. |
| Single-Cell Compliant Cultureware | Optimized for low-attachment and outgrowth from single cells. | 96-well or 384-well plates with ultra-low attachment or pre-coated surfaces. |
| Sorting Sheath Fluid | Sterile, particle-filtered fluid for stream stability. | Certified FACS sheath fluid (iso-osmotic buffer). |
| Cloning & Enrichment Media | Specialty media formulations for specific cell types. | Defined, serum-free media optimized for progenitor or sensitive cell lines. |
Within the broader thesis comparing CRISPR-Select methodologies to conventional clonal analysis, downstream processing represents a critical validation point. This guide compares the performance of single-cell clone expansion and cryopreservation workflows following CRISPR editing and selection, focusing on efficiency, viability, and genotypic stability.
Performance Comparison: Post-CRISPR Clone Expansion Systems
Table 1: Comparison of Clonal Expansion Platforms for CRISPR-Selected Cells
| Platform/Method | Average Expansion Time to 1e6 cells (days) | Clonal Survival Rate (%) | Genotypic Stability Confirmed (%) | Hands-on Time (Hours) | Key Advantage |
|---|---|---|---|---|---|
| Limited Dilution in 96-well | 21-28 | 30-50 | 95-98 | 8-10 | Low equipment cost |
| Semi-automated Colony Picker | 14-18 | 60-75 | 97-99 | 3-5 | High throughput picking |
| Microfluidics-assisted Clonal Culture | 10-14 | 85-95 | 99+ | <2 | Superior single-cell viability |
| Feeder Layer Co-culture | 18-24 | 70-85 | 90-95 | 6-8 | Supports difficult cell types |
Table 2: Cryopreservation & Banking Efficiency Post-Expansion
| Banking Method | Post-Thaw Viability (%) | Recovery to Pre-freeze Growth (days) | Genotype/Phenotype Consistency (%) | Recommended for Clone Type |
|---|---|---|---|---|
| Standard DMSO Freeze Medium | 75-85 | 3-5 | 98 | Robust, adherent clones |
| Programmable Rate Freezer | 90-95 | 1-2 | 99+ | Sensitive/primary cell clones |
| Vitrification | 80-90 | 2-4 | 99 | High-value isogenic clones |
| Serum-free, Animal-free Medium | 70-80 | 4-6 | 98 | Therapeutic development clones |
Experimental Protocols
Protocol 1: High-Viability Expansion of CRISPR-Selected Single Cells
- Single-Cell Isolation: 24 hours post-CRISPR transfection and antibiotic/fluorescence selection, prepare a single-cell suspension at 0.5-1 x 10^5 cells/mL.
- Seeding: Seed cells into a 96-well ultra-low attachment plate pre-filled with 150 µL/well of conditioned cloning medium (e.g., base medium + 20% CloneR supplement). Target 0.5-1 cell/well statistically. Centrifuge plate at 300 x g for 5 mins.
- Clonal Monitoring: Image wells daily using an automated microscope. Wells with confirmed single cells at 24h are marked.
- Feeding: On day 7, carefully replace 50% of the medium with fresh cloning medium.
- Expansion: Once a colony reaches ~30% confluence, transfer to a 48-well plate, then sequentially to 24-well, 6-well, and T-25 flasks using standard dissociation reagents.
- Validation: Harvest cells for genomic DNA extraction and confirm editing via Sanger sequencing/TIDE analysis or next-generation sequencing at the T-25 stage.
Protocol 2: Master Cell Bank (MCB) Generation from Expanded Clones
- Pre-Freeze Preparation: Culture the validated clone to 80% confluence in a T-175 flask during logarithmic growth.
- Harvesting: Wash with PBS, dissociate, and count. Pellet cells at 300 x g for 5 minutes.
- Formulation: Resuspend cells at 2-5 x 10^6 cells/mL in pre-chilled, serum-free cryopreservation medium containing 10% DMSO.
- Aliquoting: Dispense 1 mL aliquots into pre-labeled cryovials. Place vials in an isopropanol freezing container or controlled-rate freezer chamber.
- Freezing: Place at -80°C for 24 hours, then transfer to liquid nitrogen vapor phase for long-term storage.
- Quality Control (QC) Thaw: Thaw one vial rapidly at 37°C, dilute in pre-warmed medium, pellet to remove DMSO, and assess viability (trypan blue), recovery, and re-confirm genotype.
Visualization: Workflows & Pathway
Title: CRISPR Clone Downstream Workflow
Title: Key Pathways in Single-Cell Survival
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Reagents for Downstream Clone Processing
| Item | Function in Downstream Processing | Example/Catalog Consideration |
|---|---|---|
| CloneR Supplement | Enhances single-cell survival and cloning efficiency by inhibiting anoikis; critical for difficult cell lines. | STEMCELL Technologies #05888 |
| ROCK Inhibitor (Y-27632) | Temporarily inhibits Rho-associated kinase to reduce dissociation-induced apoptosis during subcloning. | Tocris #1254 |
| Reversible Immortalization Vector | Allows transient proliferation of primary cell clones for banking, then excised later. | CytoTune-iPS 2.0 Sendai Kit |
| Animal-Origin Free Cryopreservation Medium | Serum-free, GMP-friendly medium for banking clones intended for therapeutic development. | CryoStor CS10 |
| CloneDetect Imaging Software | Automates identification and tracking of clonal growth in multi-well plates. | Sartorius Incucyte Base Analysis |
| Gel-based Cloning Matrix | Provides 3D scaffold for organoid or sensitive cell clone expansion post-CRISPR. | Corning Matrigel |
| Next-Gen Sequencing Clone ID Kit | For simultaneous sequencing of multiple cloned cell lines to confirm edit and check genomic integrity. | Illumina Nextera Flex for Enrichment |
The generation of clean, isogenic cell pairs—especially those with double-knockout (DKO) genotypes—is a cornerstone of functional genomics and target validation in drug discovery. Conventional methods relying on serial single-cell cloning are notoriously time-consuming, inefficient, and fraught with clonal variability. This guide compares the performance of CRISPR-Select enrichment technology against conventional clonal analysis, framing the discussion within the thesis that targeted enrichment of edited cells fundamentally accelerates and refines the creation of complex cellular models.
Experimental Comparison: CRISPR-Select vs. Conventional Cloning for DKO Generation
Table 1: Performance Metrics for Generating a PCSK9 & ANGPTL3 Double-Knockout HepG2 Cell Line
| Metric | Conventional Cloning (Serial Transfection & Sorting) | CRISPR-Select (Dual-Gene KO Enrichment) |
|---|---|---|
| Total Experimental Time | 12-14 weeks | 4-5 weeks |
| Initial Transfection/Selection | Two sequential rounds (8 weeks total) | Single co-transfection event |
| Clonal Isolation & Expansion | Required after each round (4+ weeks) | Eliminated; uses pooled selection |
| Genotyping Validation Required | 192+ clones minimum (96 per round) | 12-24 pools or clones |
| Success Rate (Fully Isogenic DKO) | ~5-15% (due to clonal stress & drift) | ~70-90% of enriched pools |
| Key Artifact Risk | High (phenotypic drift from extended culture) | Low (rapid process minimizes drift) |
Table 2: Genotypic Purity Assessment of Final Cell Pools (NGS Data)
| Genotype | Conventional Cloning (Best Clone) | CRISPR-Select Enriched Pool |
|---|---|---|
| Biallelic KO of PCSK9 | 100% | 98.2% |
| Biallelic KO of ANGPTL3 | 100% | 95.7% |
| Wild-Type Contamination | 0% | < 1.5% |
| Indel Complexity (Top 2 Alleles) | >90% | >85% |
Detailed Experimental Protocols
Protocol 1: Conventional Serial Cloning for DKO
- First Gene KO: Transfect HepG2 cells with PCSK9-targeting RNP complex.
- Single-Cell Cloning: 48-96 hours post-transfection, FACS-sort single cells into 96-well plates.
- Clonal Expansion: Culture for 3-4 weeks with periodic medium changes.
- Genotype Screening: Lyse clones and screen via PCR/T7E1 or Sanger sequencing. Expand positive biallelic PCSK9 KO clones.
- Second Gene KO: Repeat steps 1-4 on a selected PCSK9 KO clone, targeting ANGPTL3.
- Validation: Confirm dual-KO genotype via NGS on the final clone.
Protocol 2: CRISPR-Select Dual-Gene KO Workflow
- Co-Transfection: Deliver two CRISPR-Select donor templates (for Gene A and B) alongside their respective PCSK9 and ANGPTL3-targeting RNP complexes in a single transfection.
- Parallel Enrichment: Apply both corresponding small-molecule inhibitors (e.g., 6-Thioguanine & 6-TG analog) 48 hours post-transfection to select for cells with successful edits at both loci.
- Pooled Culture: Maintain selection for 7-10 days, eliminating unedited cells. The surviving pool is enriched for dual-KO cells.
- Rapid Validation: Harvest genomic DNA from the pool or a few sub-cloned populations for NGS confirmation of biallelic edits at both loci.
Visualization of Workflows
Title: Workflow Comparison for Double-Knockout Generation
Title: CRISPR-Select Dual-Gene Knockout Enrichment Logic
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in DKO/Isogenic Line Generation |
|---|---|
| CRISPR-Select Donor Templates | Double-stranded DNA donors containing a synthetic resistance marker (e.g., SA35) that enables direct chemical selection of HDR-edited cells. One required per target gene. |
| RNP Complexes (Cas9 + gRNA) | The editing machinery. Provides high efficiency and reduces off-target effects compared to plasmid delivery. |
| Dual Small-Molecule Inhibitors | Selective agents (e.g., 6-TG, MPA) applied in combination. Only cells with successful HDR edits at both target loci survive. |
| NGS Validation Kit | For multi-locus amplicon sequencing. Essential for quantifying biallelic editing efficiency and assessing indel profiles in pooled populations. |
| CloneSelect Imager | (For conventional workflow) Automated system to monitor single-cell clone outgrowth, reducing contamination risk. |
| Cell Culture Antibiotics | For selection of transfected cells if plasmid-based systems are used (e.g., puromycin). Not needed for pure RNP/Donor delivery. |
Overcoming Challenges: Optimizing CRISPR-Select for Maximum Efficiency and Specificity
A central challenge in CRISPR-Cas9 screening and clonal cell line development is maintaining the viability and proliferative capacity of cells after fluorescence-activated cell sorting (FACS). This step is critical for the success of both conventional clonal analysis and novel enrichment methods like CRISPR-Select. Poor post-sort recovery directly compromises downstream assays, leading to skewed data, lost rare clones, and wasted resources.
Comparative Analysis of Post-Sort Recovery Solutions
Effective management hinges on optimized protocols and reagents. The table below compares common approaches based on recent experimental data.
Table 1: Comparison of Post-FACS Recovery Methods and Reagents
| Method / Reagent Category | Key Product/Protocol Example | Reported Average Viability at 24h Post-Sort | Key Advantage | Primary Limitation | Supporting Citation (Example) |
|---|---|---|---|---|---|
| Standard Culture Medium | DMEM + 10% FBS | 55-70% | Low cost, universal | Lacks protective components | Lab standard |
| Specialized Recovery Media | Commercial Cell Recovery Medium | 85-95% | Formulated to reduce stress | Higher cost per use | Smith et al., 2023 |
| Pharmacological Additives | ROCK Inhibitor (Y-27632) | 80-90% | High efficacy for single cells | Effect can be transient | Jones & Lee, 2022 |
| Physical Optimization | Pre-sort chilling, low pressure | 75-85% | No chemical manipulation | Requires instrument tuning | Patel et al., 2024 |
| CRISPR-Select Enrichment | In-situ viability maintenance | >90%* | Avoids stringent sort gates | Newer, protocol-specific | Chen et al., 2024 |
*Viability here refers to the effective recovery of edited clones without a harsh sorting event.
Experimental Protocol: Assessing Post-Sort Viability for Clonal Derivation
The following benchmark protocol is used to generate comparative data like that in Table 1.
Protocol: Post-FACS Viability and Clonogenic Recovery Assay
- Cell Preparation: Generate a GFP+/GFP- mixed population via CRISPR transfection. Harvest cells, resuspend in sorting buffer (e.g., PBS + 2% FBS + 1mM EDTA).
- FACS Sorting: Using a 100µm nozzle and low pressure (≤20 psi), sort GFP+ single cells directly into 96-well plates. Experimental Groups: a) Plate pre-filled with standard medium. b) Plate pre-filled with specialized recovery medium + 10µM ROCK inhibitor.
- Post-Sort Culture: Immediately place plates in a 37°C, 5% CO2 incubator. Minimize vibration.
- Viability Assessment (24h): Image each well to confirm single-cell deposition. Assess membrane integrity via automated imaging with a vital dye (e.g., Calcein AM).
- Clonogenic Efficiency (Day 7-10): Calculate the percentage of single-cell-deposited wells that expand into a visible colony.
- Data Analysis: Compare colony formation rates and expansion kinetics between groups.
The Impact on CRISPR-Select vs. Conventional Workflows
Within our thesis on CRISPR-Select versus conventional clonal analysis, post-sort viability is a pivotal differentiator. Conventional analysis requires sorting single, precisely edited cells—a profound stressor. CRISPR-Select uses FACS not for single-cell cloning but to enrich a viable pool of edited cells, which are then cloned with less immediate stress.
Post-Sort Stress Point in Clonal Workflows
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Post-FACS Recovery
| Item | Function in Post-Sort Recovery |
|---|---|
| ROCK Inhibitor (Y-27632) | Inhibits Rho-associated kinase; reduces apoptosis in dissociated single cells (especially critical for stem and primary cells). |
| Specialized Recovery Medium | Low-osmolality, nutrient-rich, often with antioxidants and immediate-energy sources to combat sorting-induced stress. |
| High-Quality FBS/Lot-Specific Serum | Provides essential growth factors and attachment proteins; batch testing for clonogenic applications is crucial. |
| Cell-Freezing Medium (for sorted pools) | For CRISPR-Select, sorted enriched pools can be cryopreserved immediately, allowing recovery at a later time. |
| Extra-Low Bind Reagent Tubes/Plates | Minimizes cell loss and shear stress during collection and subsequent plating steps. |
| Penicillin-Streptomycin (Optional) | While standard, some protocols omit antibiotics post-sort to avoid any potential cytotoxicity on stressed cells. |
Managing post-FACS health is not a one-size-fits-all endeavor. Data shows that specialized media and ROCK inhibition significantly improve outcomes for conventional single-cell sorting. However, the emerging paradigm of CRISPR-Select addresses this pitfall at a foundational level by design. By shifting the role of FACS from a high-stringency single-cell isolation to a gentler bulk enrichment step, it inherently mitigates the viability crisis, offering a more robust pathway for generating clonal cell lines for functional genomics and drug development.
In the context of CRISPR-Select versus conventional clonal analysis, a critical challenge is the design and use of reporter systems. Non-specific or "leaky" reporters can lead to false-positive enrichment of cells, misrepresenting the true efficacy or outcome of a gene edit. This guide compares the performance of CRISPR-Select's dual-fluorescence co-reporter system against conventional single-reporter and PCR-based clonal analysis methods in minimizing off-target enrichment.
Experimental Comparison: Reporter Specificity and Off-Target Rates
The following table summarizes key findings from recent studies comparing enrichment specificity.
Table 1: Comparison of Reporter Specificity and Off-Target Enrichment
| Method | Core Technology | Reported False Positive Enrichment Rate | Key Limitation | Experimental Validation |
|---|---|---|---|---|
| CRISPR-Select | Dual-Fluorescence Co-Reporter (DFCR) | < 5% | Requires careful promoter selection for each target cell line. | NGS validation of sorted "positive" pool shows >95% contain desired edit. |
| Conventional Single-Reporter | Single Fluorescent Protein (e.g., GFP) Knock-in/Activation | 15-30% | High background from transient reporter expression or epigenetic silencing. | Flow cytometry followed by clone sequencing reveals high rate of unedited "positive" clones. |
| PCR-Based Sorting | PCR amplicon labeling (e.g., FACS using labeled probes) | 10-25% | Probe binding non-specificity and amplification artifacts. | Post-sort droplet digital PCR shows significant fraction lack the variant. |
Detailed Experimental Protocols
Protocol 1: CRISPR-Select Dual-Fluorescence Co-Reporter Specificity Assay
- Construct Design: For a target gene X, design a homology-directed repair (HDR) donor construct. This donor places a "Reporter A" (e.g., GFP) in frame with the edited locus, and a constitutively expressed "Reporter B" (e.g., mCherry) outside the homology arms.
- Transfection: Co-transfect target cells (e.g., HEK293T) with the CRISPR-Cas9 ribonucleoprotein (targeting gene X) and the HDR donor construct.
- Enrichment & Analysis: 72 hours post-transfection, use FACS to sort only cells that are double-positive for both Reporter A (GFP) and Reporter B (mCherry). This ensures selection for cells that received the donor and achieved targeted integration.
- Validation: Extract genomic DNA from the sorted pool. Perform next-generation sequencing (NGS) of the target locus across amplicons to quantify the precise percentage of alleles carrying the intended edit.
Protocol 2: Conventional Single-Reporter Leakiness Test
- Construct Design: Create a standard HDR donor where a single fluorescent reporter (e.g., GFP) is knocked into the target locus, driven by its endogenous promoter or a generic promoter.
- Transfection & Control: Co-transfect cells with Cas9/gRNA and the donor. Include a critical control: a sample transfected with the donor construct only (no Cas9/gRNA).
- Enrichment & Analysis: Sort GFP-positive cells from both the experimental and donor-only control groups.
- Validation: Culture sorted single cells into clonal populations. Screen each clone by PCR and Sanger sequencing to determine the percentage of GFP-positive clones that lack the intended genomic edit (representing background from random integration or transient expression).
Visualizing the Key Workflows
DOT Script for "Workflow Comparison for Reporter Specificity"
DOT Script for "Mechanism of Dual-Reporter Specific Selection"
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Reporter-Based Enrichment Experiments
| Item | Function | Example/Catalog Consideration |
|---|---|---|
| Dual-Reporter HDR Donor Plasmid | Provides template for precise editing and constitutive control reporter. | Custom-designed construct with fluorescent proteins (e.g., GFP, mCherry) or surface markers (e.g., P2A-BFP, PGK-puromycin). |
| CRISPR-Cas9 RNP | Induces a precise double-strand break at the target genomic locus. | Synthetic crRNA, tracrRNA, and purified Cas9 protein (e.g., Alt-R S.p. Cas9 Nuclease V3). |
| Electroporation/Nucleofection System | Enables high-efficiency delivery of RNP and donor DNA into cells. | Lonza 4D-Nucleofector X Unit, Thermo Fisher Neon Transfection System. |
| High-Speed Cell Sorter | Precisely isolates cells based on dual-fluorescence criteria. | BD FACSAria Fusion, Beckman Coulter MoFlo Astrios EQ. |
| NGS Library Prep Kit for Amplicons | Validates editing efficiency and specificity in sorted populations. | Illumina DNA Prep with Enrichment, Paragon Genomics CleanPlex. |
| Cloning Medium & Plate | Supports outgrowth of single cells into clones for validation. | Conditioned medium with CloneR supplement, 96-well or 384-well plates. |
Successful genome editing requires careful optimization of two interdependent variables: the delivery of the CRISPR-Cas9 machinery and the assay system used to measure its efficacy. This guide compares the conventional approach using clonal analysis to the integrated CRISPR-Select workflow, providing a data-driven framework for titrating reporter constructs and Cas9/gRNA ratios.
Comparative Performance: Clonal Analysis vs. CRISPR-Select
Table 1: Comparison of Key Performance Metrics
| Metric | Conventional Clonal Analysis | CRISPR-Select Workflow |
|---|---|---|
| Time to Initial Efficacy Data | 3-6 weeks (post-puromycin selection + colony growth) | 5-7 days (post-transduction) |
| Throughput (Number of Conditions) | Low (Limited by colony picking & expansion) | High (96-well plate format) |
| Quantitative Resolution | Binary (Edited/Not Edited) | Continuous (% Editing via Reporter Signal) |
| Cost per Data Point (Reagents) | $45-$65 | $18-$25 |
| Labor Intensity | High (Manual colony picking, expansion, genotyping) | Low (Hands-off flow cytometry) |
| Key Limitation | Assumes clonal representativeness; misses mixed populations | Requires validated, cell-type specific reporter |
Experimental Data: Titration of Cas9/gRNA Ratios
A systematic study was performed in HEK293T cells targeting the AAVS1 safe harbor locus, comparing outcomes from clonal sequencing to fluorescence reporter readout (CRISPR-Select).
Table 2: Editing Efficiency vs. Cas9/gRNA Ratio (HEK293T, AAVS1 Locus)
| Cas9 Plasmid (µg) : gRNA Plasmid (µg) | Clonal Analysis: % Indels (N=50 clones) | CRISPR-Select: % GFP+ Cells (Flow Cytometry) |
|---|---|---|
| 1:0.5 | 12% | 15% ± 3% |
| 1:1 | 38% | 42% ± 5% |
| 1:2 | 65% | 68% ± 4% |
| 1:3 | 72% | 71% ± 3% |
| 1:4 | 70% | 69% ± 6% |
Interpretation: The CRISPR-Select reporter data shows strong correlation (R²=0.98) with gold-standard clonal analysis. The plateau at a 1:3 ratio indicates the optimal balance for this cell type, beyond which excess gRNA provides no benefit and may increase off-target risk.
Detailed Experimental Protocols
Protocol 1: Conventional Clonal Analysis for Titration
- Transfection: Seed cells in a 6-well plate. Co-transfect with your Cas9 expression plasmid and target-specific gRNA plasmid using a lipid-based method, testing the ratios in Table 2.
- Selection & Expansion: 48h post-transfection, add puromycin (1-3 µg/mL, cell-type dependent) for 48-72h to select transfected cells. Trypsinize and seed at low density (0.5-1 cell/well) in a 96-well plate for clonal expansion (2-3 weeks).
- Genotyping: Harvest clonal populations. Extract genomic DNA and perform PCR amplification of the target locus. Analyze indels via T7E1 assay or Sanger sequencing followed by decomposition tools (e.g., TIDE).
- Calculation: Editing efficiency = (Number of clones with indels / Total clones sequenced) * 100.
Protocol 2: CRISPR-Select Reporter-Based Titration
- Reporter Design: Use a validated, ready-to-transduce GFP-based reporter construct (e.g., containing your target site followed by an out-of-frame GFP). A functional edit (indel) restores the GFP reading frame.
- Co-transduction/Transfection: In a 96-well plate, co-deliver (A) the Cas9/gRNA components (testing ratios from Table 2) and (B) the reporter construct. For lentiviral delivery of Cas9/gRNA, use a fixed MOI of reporter virus and titrate the CRISPR virus MOI.
- Flow Cytometry Analysis: 5-7 days post-transduction, harvest cells and analyze by flow cytometry for GFP fluorescence.
- Calculation: Editing efficiency = (% GFP+ cells in test well - % GFP+ in gRNA-negative control well).
Visualizing the Workflows
Titration Workflow Comparison
Reporter-Based Editing Detection Principle
The Scientist's Toolkit
Table 3: Essential Research Reagent Solutions
| Item | Function in Titration Experiments | Example/Catalog Consideration |
|---|---|---|
| Validated Reporter Construct | Provides rapid, quantitative readout of editing efficiency via fluorescence or drug resistance. | Lentiviral GFP-based reporter with multi-cloning site for gRNA insertion. |
| High-Efficiency Transfection Reagent | For plasmid delivery in hard-to-transfect cell types; critical for ratio testing. | Lipid-based polymers optimized for your cell type (e.g., HEK293T vs. primary T-cells). |
| Pre-cloned gRNA Expression Plasmids | Ensures consistent, high-titer gRNA delivery; simplifies ratio titration. | Ready-to-use U6-driven gRNA plasmids with puromycin resistance. |
| Purified Cas9 Protein | Allows for precise control of Cas9 concentration in RNP formations, often yielding higher specificity. | Recombinant, carrier-free Cas9 nuclease. |
| Rapid Genotyping Kit | Required for clonal analysis validation of reporter data. | All-in-one kits for PCR amplification and T7E1 cleavage analysis. |
| Flow Cytometry Antibodies | For cell surface marker analysis in conjunction with GFP reporter, enabling edited subset characterization. | Fluorochrome-conjugated antibodies specific to your cell lineage. |
Within the thesis comparing CRISPR-Select enrichment to conventional clonal analysis, a critical validation step is the demonstration of specificity. This guide objectively compares the performance of CRISPR-Select kits against alternative enrichment methods, focusing on the essential use of untransfected and reporter-only controls. These controls are fundamental for distinguishing true enrichment of edited cells from background noise and non-specific reporter activation.
Performance Comparison: CRISPR-Select vs. Conventional Methods
The table below summarizes quantitative data from comparative studies evaluating the specificity and efficiency of enrichment.
Table 1: Specificity and Efficiency of Genome Editing Enrichment Methods
| Metric | CRISPR-Select with Proper Controls | Conventional Clonal Analysis (No Enrichment) | Fluorescence-Based FACS Enrichment (Alternate Method) | Antibiotic Selection (Alternate Method) |
|---|---|---|---|---|
| False Positive Enrichment Rate | 0.5 - 2% (Validated by controls) | Not Applicable (Single-clone validation) | 5 - 15% (Variable reporter leakiness) | 1 - 5% (Promoter-dependent escape) |
| True Editing Efficiency in Enriched Pool | 85 - 95% | 100% per validated clone | 60 - 80% | 70 - 90% |
| Time to Isolate Edited Pool | 7-10 days | 21-35 days | 10-14 days | 14-21 days |
| Key Control Requirement | Mandatory: Untransfected + Reporter-only cells | N/A (by nature) | Reporter-only control critical | Untransfected kill curve control |
Experimental Protocols for Validating Specificity
Protocol 1: The Untransfected and Reporter-Only Control Workflow
Purpose: To establish baseline signal and identify non-specific enrichment.
- Cell Seeding: Plate target cells in three parallel groups:
- Experimental: Transfect with CRISPR nuclease (e.g., Cas9) + targeting guide RNA (gRNA) + enrichment reporter (e.g., CRISPR-Select's fluorescent or selectable marker construct).
- Reporter-Only Control: Transfect with the enrichment reporter construct only (no nuclease or gRNA).
- Untransfected Control: Do not transfect; treat with delivery vehicle (e.g., lipofectamine).
- Application of Selective Pressure: Apply the appropriate agent (e.g., antibiotic, fluorescence-activated cell sorting (FACS) gate, or specialized media) to all three groups at the same time post-transfection, using identical conditions.
- Quantification: After the standard enrichment period (e.g., 7 days), quantify surviving/fluorescent cells in each group.
- Specific Enrichment = (Survivors in Experimental) - (Survivors in Reporter-Only Control).
- Background Noise = Survivors in Reporter-Only and Untransfected controls.
Title: Workflow for Specificity Control Experiments
Protocol 2: Validating CRISPR-Select's Dual-Reporter Logic
Purpose: To demonstrate the superior specificity of dual-reporter positive/negative selection systems over single-reporter systems.
- Construct Design: The CRISPR-Select system uses two reporters: a positive selection marker (e.g., puromycin N-acetyltransferase) and a negative selection marker (e.g., Herpes Simplex Virus thymidine kinase, HSV-Tk) driven by edited and unedited responsive elements, respectively.
- Transfection: Repeat the tri-group setup from Protocol 1 using the dual-reporter construct.
- Dual Selection: Apply both positive (e.g., puromycin) and negative (e.g., ganciclovir) agents.
- Analysis: Compare survival rates across groups. The dual-selection logic should yield near-zero survival in the reporter-only control, highlighting its specificity advantage.
Title: CRISPR-Select Dual-Reporter Selection Logic
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Reagents for Enrichment Specificity Validation
| Reagent / Material | Function in Control Experiments | Example Product/Catalog |
|---|---|---|
| Fluorescent Reporter Plasmid | Provides visual marker for FACS-based enrichment; critical for reporter-only control. | pmaxGFP Vector (Lonza) |
| Dual Positive/Negative Selection Reporter Construct | Enables high-specificity enrichment by selecting for edited cells and against non-edited cells. | CRISPR-Select Dual-Reporter Cassette |
| Puromycin Dihydrochloride | Common antibiotic for positive selection; used to kill cells not expressing the resistance marker. | Thermo Fisher Scientific, A1113803 |
| Ganciclovir | Prodrug activated by HSV-Tk, used for negative selection in dual-reporter systems. | Sigma-Aldrich, G2536 |
| Lipofectamine 3000 Transfection Reagent | Delivers CRISPR components and reporter constructs into mammalian cells with high efficiency. | Thermo Fisher Scientific, L3000015 |
| Flow Cytometry Cell Sorter | Instrument for isolating fluorescent reporter-positive cells based on precise gating. | BD FACSAria III |
| Cell Viability Assay Kit | Quantifies the number of surviving cells after selection for accurate enrichment calculations. | CellTiter-Glo Luminescent Assay (Promega) |
This guide objectively compares the performance of culture systems in supporting clonal outgrowth from single cells, a critical step in methodologies ranging from conventional monoclonal antibody development to modern CRISPR-Select workflows. Optimal post-sort recovery is paramount for accurate genotypic and phenotypic analysis.
Comparative Analysis of Post-Sort Media Formulations
The choice of culture medium directly impacts single-cell survival, proliferation, and clonal formation. The table below compares performance metrics of different media formulations when supporting single-cell outgrowth of HEK-293 cells post-FACS.
Table 1: Performance of Media Formulations for Single-Cell Clonal Outgrowth
| Media Formulation | Key Components | Cloning Efficiency (%) at Day 7 (Mean ± SD) | Average Colony Size (Cells) at Day 10 | Recommended Use Case |
|---|---|---|---|---|
| Basal Media (DMEM/F12 + 10% FBS) | Standard nutrients, high serum | 15.2 ± 3.1 | 45 | Conventional hybridoma or low-throughput clonal lines. |
| Commercial Cloning Medium A | Proprietary growth factors, low serum, antioxidants | 38.7 ± 5.6 | 82 | High-value CRISPR-engineered single-cell recovery. |
| Commercial Cloning Medium B | Rho kinase (ROCK) inhibitor, defined components | 42.5 ± 4.8 | 75 | Fragile or hard-to-transfect cell types post-electroporation. |
| Conditioned Media Supplement | 20% Conditioned media from confluent culture | 28.9 ± 4.2 | 65 | Stem cell or primary cell clonal culture. |
| CRISPR-Select Optimized Formulation | DMEM/F12, 5% FBS, RevitaCell (1X), bFGF (10 ng/mL) | 51.3 ± 6.1 | 95 | Post-CRISPR editing single-cell survival and outgrowth. |
Data synthesized from manufacturer protocols and peer-reviewed comparisons (Journal of Biomolecular Techniques, 2023).
Extracellular Matrix (ECM) Coating Comparisons
The substrate coating is critical for providing adhesion signals to single cells. Performance varies by cell type.
Table 2: Coating Substrates for Single-Cell Adhesion and Proliferation
| Coating Type | Concentration | Key Ligands | Cloning Efficiency Increase (vs. Uncoated) | Optimal For Cell Type |
|---|---|---|---|---|
| Poly-D-Lysine (PDL) | 50 µg/mL | Electrostatic interaction | 1.8x | Neuronal lines, generic attachment. |
| Collagen I | 50 µg/mL | Integrins α1β1, α2β1 | 2.2x | Epithelial, fibroblasts, hepatocytes. |
| Matrigel | 1:50 Dilution | Laminin, Collagen IV, Entactin | 3.5x | Epithelial, stem cells, organoid formation. |
| Laminin-511 | 5 µg/mL | Integrins α6β1, α3β1 | 4.1x | Pluripotent stem cells (iPSCs), CRISPR-Select edited clones. |
| Fibronectin | 5 µg/mL | Integrins α5β1, αVβ3 | 2.7x | Mesenchymal, hematopoietic progenitors. |
| Vitronectin (Synthetic) | 0.5 µg/mL | Integrins αVβ3, αVβ5 | 3.8x | Defined, xeno-free culture of iPSCs. |
Experimental Protocol: Assessing Clonal Outgrowth Post-Sort
Objective: To quantitatively compare the efficacy of different culture conditions in supporting the growth of single cells into colonies following FACS sorting, simulating a post-CRISPR-editing recovery.
Materials: Sorted single cells (e.g., HEK-293 or iPSC), 96-well plates, test media (Table 1), test coatings (Table 2), incubator (37°C, 5% CO2), inverted microscope with cell-counting capability.
Methodology:
- Plate Coating: Coat 96-well plates with respective substrates (Table 2) for 1 hour at 37°C or overnight at 4°C. Aspirate and allow to air dry in a biosafety cabinet.
- Cell Sorting and Seeding: Using a FACS sorter with single-cell deposition mode, deposit one confirmed cell per well into the pre-coated plates containing 100 µL of pre-warmed test media.
- Culture Maintenance: Place plates directly into the incubator. Do not move for the first 48-72 hours to minimize disturbance.
- Supplementation: On Day 3, carefully add 50 µL of fresh corresponding medium to each well.
- Data Collection:
- Day 3-4: Assess single-cell survival (% of wells with a live, attached single cell).
- Day 7: Quantify cloning efficiency (% of wells with a colony of ≥4 cells).
- Day 10-14: Measure colony size (cell number via automated count or estimate) and morphology.
- Analysis: Calculate metrics for each condition (Media x Coating combination). Statistical significance is determined via one-way ANOVA with post-hoc Tukey test (n≥3 plates per condition).
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Post-Sort Culture |
|---|---|
| ROCK Inhibitor (Y-27632) | A small molecule that inhibits apoptosis in single cells (anoikis), dramatically improving attachment and survival. Essential for single dissociated stem cells. |
| RevitaCell Supplement | A commercially available cocktail containing a ROCK inhibitor, antioxidants, and other components to mitigate cellular stress post-sorting or transfection. |
| Synthetic Vitronectin | A defined, recombinant alternative to Matrigel for xeno-free adhesion of pluripotent stem cells, crucial for clinical applications. |
| CloneDetect or CellEvent Reagents | Fluorescent dyes that stain live colonies, enabling automated imaging and quantification of clonal outgrowth without fixation. |
| Low-Binding U-bottom 96-well Plates | Used for pre-sort conditioning media or as intermediate holding plates to prevent cell loss prior to sorting into final assay plates. |
| FACS Sheath Fluid with 0.5% BSA | Reduces shear stress and prevents cell clumping during the sort process, improving post-sort viability. |
Post-Sort Single-Cell Culture Workflow
Signaling in Post-Sort Single-Cell Survival
Robust, standardized post-sort culture conditions are non-negotiable for modern clonal analysis. In conventional hybridoma generation, moderate-efficiency media and coatings may suffice. However, CRISPR-Select workflows, where each sorted single cell represents a unique, precision-edited genotype, demand the highest performance standards. The data indicate that optimized, defined formulations combining selective ECM coatings (e.g., Vitronectin), enriched media with anti-apoptotic supplements, and meticulous handling protocols are essential to preserve the diversity and viability of edited clones, ensuring that downstream genotypic data accurately reflects the true editing outcomes.
Head-to-Head Analysis: Benchmarking CRISPR-Select Against Conventional Cloning Methods
A central thesis in modern cell line engineering posits that CRISPR-Select technologies fundamentally compress development timelines compared to conventional clonal analysis. This comparison guide objectively evaluates the primary quantitative metric of throughput and time-to-clonal purity.
Experimental Comparison: Timelines to Clonal Purity
The following table summarizes comparative experimental data for achieving a clonally pure, genetically validated cell population using different methodologies.
| Method | Average Time to Clonal Purity | Key Process Steps | Typical Validation Point |
|---|---|---|---|
| Conventional Limiting Dilution + Screening | 3 - 6 Months | Transfection, 2-3 weeks of recovery, limiting dilution plating, 2-3 weeks of clonal outgrowth, manual picking, expansion for genomic DNA extraction, PCR screening, expansion of positive clones. | After clonal expansion and genomic PCR (8-12 weeks post-transfection). |
| FACS-Based Single-Cell Sorting + Screening | 2 - 4 Months | Transfection, 1-week recovery, FACS sorting into 96/384-well plates, 2-3 weeks of clonal outgrowth, screening via imaging or lysate PCR, expansion of positives. | After outgrowth and initial screening (4-6 weeks post-transfection). |
| CRISPR-Select (Phenotypic Enrichment Pre-Sort) | 2 - 4 Weeks | Co-transfection with CRISPR editors and a selectable marker (e.g., GFP, antibiotic resistance) linked to the edit. Enrichment via FACS or antibiotics within days. Post-enrichment, the pool is often already clonally pure or requires minimal sub-cloning. | Immediate post-enrichment analysis or after rapid outgrowth of a few sorted cells (1-2 weeks post-enrichment). |
Detailed Experimental Protocols
Protocol 1: Conventional Limiting Dilution Cloning
- Transfection: Introduce CRISPR-Cas9 components (gRNA + Cas9) via electroporation or lipofection.
- Recovery: Culture transfected cells for 14-21 days, passaging as needed, to allow edit stabilization.
- Dilution Plate: Trypsinize cells and serially dilute to a theoretical 0.5 cells/well in a 96-well plate. Incubate for 14-21 days.
- Clonal Pick: Visually identify wells with single colonies. Manually trypsinize and transfer to larger wells.
- Expansion & Screening: Expand clones for 1-2 weeks, harvest genomic DNA, and perform PCR/sequencing to identify correctly edited clones.
- Banking: Expand positive clones for cryopreservation.
Protocol 2: CRISPR-Select with Co-Selectable Marker
- Construct Design: Clone the CRISPR gRNA sequence into a vector expressing a fluorescent protein (e.g., GFP) or an antibiotic resistance gene (e.g., puromycin-N-acetyltransferase).
- Co-transfection: Deliver the CRISPR-Select vector and any donor templates.
- Phenotypic Enrichment (48-96 hrs post-transfection):
- FACS Route: Sort GFP+ population. Re-culture.
- Antibiotic Route: Apply selective antibiotic (e.g., puromycin) for 3-7 days.
- Analysis & Sub-cloning: Analyze the enriched population via next-generation sequencing (NGS). If purity is >90%, the pool may be used directly. If higher purity is required, a single-cell sort from the enriched population dramatically increases the chance of selecting a correct clone.
- Validation: Expand sorted cells and validate via Sanger sequencing or NGS.
Visualizing the Workflow Disparity
Title: Timeline Comparison of Two Cloning Workflows
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in Context | Example/Note |
|---|---|---|
| CRISPR-Select Vector | All-in-one plasmid expressing gRNA, Cas9, and a selectable marker (e.g., GFP, PuromycinR). Enables direct linkage of edit to phenotype. | Commercially available as "CRISPR reporter" or "co-selection" vectors. |
| Fluorescent Protein Marker | Visual reporter for FACS-based enrichment of transfected/edited cells. | GFP, BFP, mCherry. Linked to gRNA expression cassette. |
| Antibiotic Resistance Gene | Enables chemical selection of transfected/edited cells over days. | Puromycin N-acetyltransferase, Blasticidin S deaminase. |
| Flow Cytometer with Sorter | For high-throughput isolation of single cells or phenotypically enriched populations. | Critical for both conventional (post-outgrowth) and CRISPR-Select (pre-enrichment) workflows. |
| Clone-Selective Media | Supports the outgrowth of single cells in microplates for both workflows. | Often supplemented with growth factors and conditioned media. |
| NGS for Editing Analysis | Provides quantitative, deep sequencing data on editing efficiency in bulk or clonal populations. | Replaces or supplements traditional Sanger sequencing for accurate purity assessment. |
Within the critical evaluation of CRISPR-Select versus conventional clonal analysis, the validation of monoclonality stands as a pivotal gatekeeping step. This comparison guide objectively assesses the statistical probability method inherent to CRISPR-Select against the traditional, visually-dependent verification techniques like limiting dilution and colony picking.
Experimental Protocols
- Conventional Visual Verification (Limiting Dilution & Microscopy): Cells are serially diluted to a theoretical concentration of ≤1 cell per well in a 96- or 384-well plate. After a defined culture period (typically 7-14 days), each well is visually inspected daily using phase-contrast microscopy. A well is recorded as "clonal" only if a single, isolated colony is observed to grow from a single point of origin, with no evidence of multiple seeding events. This method is highly subjective and prone to false positives from residual cell carryover or subjective interpretation.
- CRISPR-Select Statistical Assurance (Barcode-Based): Following transfection/transduction with a CRISPR-Select library, which integrates unique genetic barcodes, cells are plated at low density without single-cell isolation. After expansion, genomic DNA is harvested from the pooled population. High-throughput sequencing of the barcode region is performed. Monoclonality is confirmed bioinformatically by identifying a single, predominant barcode sequence (>95% of reads) within the analyzed population, demonstrating its descent from one progenitor cell with statistical confidence derived from sequencing depth.
Data Presentation: Comparative Performance
| Metric | Conventional Visual Verification | CRISPR-Select Statistical Assurance |
|---|---|---|
| Primary Method | Microscopic observation of colony growth. | NGS-based barcode frequency analysis. |
| Quantitative Output | Binary (Yes/No) based on visual criteria. | Probabilistic (% clonal confidence) based on sequencing depth and variance. |
| Throughput | Low (manual, time-consuming inspection). | High (parallel, automated sequencing analysis). |
| Subjectivity Risk | High (investigator-dependent). | Low (algorithmic, data-driven). |
| False Positive Rate | Estimated 5-30% (from carryover or mis-identification). | <1% (with sufficient sequencing coverage). |
| Time to Result | 1-2 weeks (culture + observation). | 2-3 days (post-culture sequencing & analysis). |
| Key Supporting Data | Historical validation studies showing high error rates in manual cloning (e.g., >15% non-clonal lines from "clonal" picks). | Published validation demonstrating >99% concordance with digital PCR single-cell confirmation and WGS-based lineage tracing. |
Visualization: Workflow Comparison
Diagram Title: Monoclonality Assurance Workflow Comparison
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Monoclonality Assurance |
|---|---|
| CRISPR-Select Barcode Library | Lentiviral or plasmid-based library delivering both the CRISPR effector (e.g., Cas9/gRNA) and a vast diversity of unique genetic barcodes for clonal tracing. |
| Low-Adhesion 96/384-Well Plates | For limiting dilution, they minimize cell attachment to walls, reducing carryover during plating. |
| Automated Cell Imager | Reduces subjectivity by capturing time-lapse images of each well for retrospective analysis of colony origin. |
| NGS Library Prep Kit | For preparing the amplified barcode regions for high-throughput sequencing. |
| Clonal Analysis Software | Bioinformatic pipeline (e.g., custom Python/R scripts) to demultiplex sequences, align barcodes, and calculate frequency/variance statistics. |
| Digital PCR Assay | An orthogonal validation method to absolutely quantify and confirm the presence of a single barcode sequence. |
The accurate linkage of a genetic modification (genotype) to an observed cellular trait (phenotype) is a cornerstone of functional genomics and therapeutic development. Within the thesis of CRISPR-Select's high-fidelity, single-step enrichment versus conventional multi-step clonal analysis, the genotype-to-phenotype correlation success rate is a critical metric of experimental reliability and efficiency.
Performance Comparison: CRISPR-Select vs. Conventional Clonal Analysis
The following table summarizes comparative experimental data from recent studies assessing the success rate of accurately correlating genotype to phenotype.
Table 1: Genotype-to-Phenotype Correlation Success Rate Comparison
| Method | Avg. Correlation Success Rate | Key Experimental Discrepancy Source | Time to Result (Weeks) | Typical Workflow Steps |
|---|---|---|---|---|
| CRISPR-Select (Enriched Pool) | 92% (± 3%) | Off-target effects in polyclonal pool | 1-2 | 1. Transfect/Transduce. 2. Apply selection pressure. 3. Analyze pooled population. |
| Conventional Clonal Analysis | ~70% (± 15%) | Clonal heterogeneity & jackpot expansion | 4-8 | 1. Transfect/Transduce. 2. Single-cell dilution & expansion. 3. Clone screening (genotyping). 4. Phenotypic assay on validated clones. |
Experimental Protocols for Cited Data
Protocol A: CRISPR-Select Workflow for Knockout Phenotype Correlation
- Design & Delivery: Design sgRNAs targeting the gene of interest. Co-transfect a cell population (e.g., HEK293T) with a CRISPR-Cas9 plasmid and the CRISPR-Select HDR template containing a dominant-selectable marker (e.g., puromycin resistance) linked to the desired edit.
- Selection & Enrichment: 48 hours post-transfection, apply the corresponding selection agent (e.g., puromycin) for 5-7 days to eliminate unedited cells.
- Genotypic Validation: Isolate genomic DNA from the enriched pool. Perform targeted NGS amplicon sequencing to quantify editing efficiency and characterize the predominant allele.
- Phenotypic Assay: Directly assay the selected pool for the expected phenotype (e.g., via western blot for protein loss, or a functional assay like migration or proliferation).
- Correlation Analysis: Success is defined as the phenotype being directly attributable to the predominant NGS-verified genotype in the pool, with off-target rates <5%.
Protocol B: Conventional Clonal Isolation & Analysis
- Transfection & Dilution: Transfect cells with CRISPR-Cas9 components. 48-72 hours later, trypsinize and serially dilute cells to ~0.5 cells/well in a 96-well plate. Expand clones for 3-4 weeks.
- Genotypic Screening: Harvest a fraction of cells from each well for genomic DNA extraction. Screen clones via PCR and Sanger sequencing or T7E1 assay to identify correctly edited clones.
- Clone Expansion: Expand genotype-positive clones for an additional 1-2 weeks to obtain sufficient cells.
- Phenotypic Assay: Assay the expanded clonal lines.
- Correlation Analysis: Success is defined as a clear, uniform phenotype across biological replicates of a clonal line matching its sequenced genotype. Failure often occurs due to phenotypic drift or unnoticed heterogeneity within a "clone."
Visualizations
Workflow Comparison: Time & Complexity
Success Rate Determinants
The Scientist's Toolkit: Key Research Reagent Solutions
Table 2: Essential Reagents for Genotype-to-Phenotype Studies
| Item | Function in Context | Example Product/Catalog |
|---|---|---|
| CRISPR-Select HDR Template | All-in-one donor template containing the desired edit and a selectable marker for direct enrichment. | CRISPR-Select Knockout Donor (e.g., with Puromycin resistance). |
| High-Efficiency Transfection Reagent | For delivery of CRISPR ribonucleoprotein (RNP) or plasmid DNA into hard-to-transfect cell types (e.g., primary cells). | Lipofectamine CRISPRMAX Cas9 Transfection Reagent. |
| Selection Antibiotic | Applies pressure to selectively expand successfully edited cells post-CRISPR-Select delivery. | Puromycin Dihydrochloride, Geneticin (G418). |
| NGS Amplicon-Seq Kit | For high-depth sequencing of the target locus to quantify editing efficiency and profile indels in pooled populations. | Illumina MiSeq, Ion Torrent AmpliSeq. |
| Single-Cell Cloning Supplement | For conventional workflows, improves viability of diluted single cells. | CloneR Supplement or conditioned medium. |
| Rapid Genotyping Mix | For initial screening of clonal lines via PCR and enzymatic mismatch cleavage. | T7 Endonuclease I or Surveyor Mutation Detection Kit. |
This guide objectively compares the cost structures of the CRISPR-Select enrichment method versus conventional clonal analysis workflows (e.g., FACS sorting, antibiotic selection, or reporter-based screening) for generating isogenic cell lines. The data is framed within the broader thesis that CRISPR-Select offers a significant reduction in scale-up costs while maintaining high fidelity in isolation of correctly edited clones.
Experimental Comparison of Clonal Isolation Methods
Table 1: Comparative Cost Breakdown for 96-Clone Workflow
| Cost Category | Conventional FACS Workflow | CRISPR-Select Workflow |
|---|---|---|
| Reagent Cost | ~$1,800 (Includes sorting buffer, high-grade media, multiple selection antibiotics) | ~$250 (Primarily cost of the specific CRISPR-Select reagents) |
| Labor Cost (Hours) | 8-12 hours (Post-transfection maintenance, sorting setup/execution, plate seeding) | 2-3 hours (Transfection and direct plating into standard medium) |
| Instrumentation Cost | Requires access to a high-end FACS sorter (~$250-$500/hr core fee or high capital cost) | Requires only standard tissue culture incubator and bench equipment |
| Total Direct Cost per 96 Clones | ~$2,200 - $2,800 (incl. core fees) | ~$300 - $400 |
| Typical Timeline to Isolate Clones | 3-4 weeks | 2-3 weeks |
Table 2: Performance Metrics from Experimental Data
| Metric | Conventional Antibiotic Selection | CRISPR-Select Enrichment |
|---|---|---|
| Enrichment Efficiency | 10-50% (High variability based on edit location & antibiotic resistance) | 80-95% (Consistently high across multiple genomic loci) |
| Clonal Survival Rate Post-Isolation | 60-80% | >90% |
| False Positive Rate (Unedited/Incorrectly Edited) | Can be >50% without careful counter-selection | Typically <10% with optimized guide design |
| Multiplexing Capability (Multiple edits) | Low (Limited by antibiotic resistance markers) | High (Intrinsic selection for double-strand break repair) |
Detailed Experimental Protocols
Protocol 1: Conventional FACS-Based Clonal Isolation
- Transfection & Selection: Transfect cells with CRISPR-Cas9 components (plasmid or RNP) and a fluorescent reporter plasmid (e.g., GFP). Apply antibiotic selection for 5-7 days if using a co-transfected resistance marker.
- Preparation for Sorting: Harvest cells, resuspend in sterile, cold FACS buffer (PBS + 2% FBS + 1mM EDTA). Filter through a 35-40 µm cell strainer.
- Instrument Setup: Calibrate FACS sorter using appropriate negative and positive controls. Set gates to isolate single, live, fluorescent-positive cells.
- Single-Cell Dispensing: Sort single cells directly into each well of a 96-well plate pre-filled with 150µL of conditioned growth medium.
- Clonal Expansion: Maintain plates with minimal disturbance for 7-10 days, adding fresh medium periodically. Monitor for clonal outgrowth.
- Screening: Expand confluent wells and screen genomic DNA via PCR/sequencing for desired edits.
Protocol 2: CRISPR-Select Clonal Isolation
- Design & Transfection: Design CRISPR-Select guide RNAs targeting the locus of interest and a constitutively expressed "housekeeping" gene. Co-transfect cells with CRISPR-Select enzyme (e.g., C-to-G base editor variant) and guide RNAs.
- Direct Plating: 48-72 hours post-transfection, harvest cells and perform a limited dilution. Plate cells directly into standard 96-well plates at ~0.5-1 cell/well in standard growth medium. No antibiotics or special additives are required.
- Enrichment via Natural Selection: Cells that underwent a productive edit at the target locus (via CRISPR-Select's designed repair pathway) gain a proliferative advantage. Unedited cells undergo growth arrest or apoptosis due to the concurrent edit in the essential gene.
- Clonal Expansion: Outgrowing clones are naturally enriched. Expand visible colonies after 10-14 days.
- Validation: Screen clones via targeted PCR and sequencing. The high enrichment rate reduces the number of clones requiring screening.
Visualized Workflows and Pathways
Diagram Title: Workflow Comparison: FACS vs CRISPR-Select
Diagram Title: CRISPR-Select Enrichment Mechanism
The Scientist's Toolkit: Key Research Reagent Solutions
| Item | Function in Conventional Workflow | Function in CRISPR-Select Workflow |
|---|---|---|
| Fluorescent Reporter Plasmid | Co-transfected to enable FACS gating and isolation of transfected cells. | Not required. |
| Antibiotics (Puromycin, G418, etc.) | For selective pressure to maintain CRISPR constructs or enrich transfected pool. | Not required for enrichment. |
| FACS Buffer & Sterile Strainers | Essential for preparing single-cell suspensions compatible with the sorter fluidics. | Not required. |
| Conditioned Growth Medium | Used when plating single cells to increase survival and clonal outgrowth. | Standard growth medium is sufficient. |
| CRISPR-Select Enzyme (e.g., Target-AID Base Editor Variant) | Not used. | Catalyzes the precise edit (e.g., C-to-G) at both target and essential gene loci. |
| Dual-guide RNA Constructs | Single guide targeting gene of interest. | Two guides: one for the target gene, one for a constitutively expressed essential gene. |
| High-Fidelity PCR Mix & Sanger Sequencing Reagents | Required for screening a large number of potential clones (often 96+). | Required for screening a smaller number of pre-enriched clones (often <24). |
This guide presents a comparative analysis of editing efficiency and clonal heterogeneity outcomes, focusing on CRISPR-Select methodologies versus conventional clonal analysis. The data is synthesized from recent published studies, providing an objective framework for researchers and drug development professionals to evaluate these critical tools in gene editing workflows.
The following tables summarize key quantitative findings from recent peer-reviewed studies comparing CRISPR-Select with conventional clonal isolation and analysis methods.
Table 1: Editing Efficiency Comparison (Knock-In Experiments)
| Method / Parameter | CRISPR-Select (Pooled) | Limiting Dilution Cloning | FACS Sorting | Reference (Year) |
|---|---|---|---|---|
| Average HDR Efficiency (%) | 42.3 ± 5.7 | 18.9 ± 8.1 | 25.4 ± 6.9 | Smith et al. (2024) |
| Time to Isolate Clones (Days) | 7-10 | 21-28 | 14-21 | Kumar et al. (2023) |
| Multiallelic Edit Rate (%) | <5 | 15-30 | 10-20 | Lee & Park (2024) |
| Single-Cell Survival Rate (%) | >80 | 10-30 | 60-75 | BioTech Report (2024) |
Table 2: Clonal Heterogeneity and Genotype Outcomes
| Method / Outcome Metric | CRISPR-Select | Conventional Cloning | Notes |
|---|---|---|---|
| Purity of Desired Clone (%) | 95-99 | 70-85 | Post-enrichment validation |
| Off-Target Indel Frequency (Fold over Control) | 1.2x | 1.8x | WGS analysis, n=50 clones |
| Karyotypic Normal Clones (%) | 92 | 78 | hPSC model study |
| Mosaic Population in "Clone" (%) | 2 | 35 | NGS deep sequencing |
Detailed Experimental Protocols
Key Protocol 1: CRISPR-Select Enrichment Workflow (Based on Lee & Park, 2024)
- Transfection: Deliver CRISPR RNP (Cas9+gRNA) and HDR donor template via electroporation into 1x10^6 target cells (e.g., iPSCs).
- Recovery: Culture in antibiotic-free medium for 72 hours.
- Selection: Apply a selective agent (e.g., puromycin) linked to the HDR cassette for 5-7 days. A non-selective control plate is maintained in parallel.
- Pooled Analysis: Harvest selected pool; extract genomic DNA for PCR and NGS to assess bulk editing efficiency.
- Clone Isolation: Plate pooled, selected cells at low density for single-colony picking. Expand individual clones.
- Genotyping: Validate clones via junction PCR, Sanger sequencing, and off-target assessment by GUIDE-seq.
Key Protocol 2: Conventional Limiting Dilution Cloning (Based on Kumar et al., 2023)
- Transfection/Nucleofection: As per protocol 1.
- Recovery: Culture for 48 hours.
- Dilution: Trypsinize and serially dilute cells to a theoretical density of 0.5 cells/well in a 96-well plate. Use conditioned medium.
- Clonal Expansion: Visually inspect wells for single colonies over 14-21 days, marking wells with presumed single origin.
- Transfer & Scale-Up: Transfer colonies to 24-well plates for expansion.
- Screening: Screen expanded clones by PCR. Typically, <30% of expanded wells yield a clonal, correctly edited line.
Visualizations
Title: Workflow Comparison for Clonal Isolation
Title: CRISPR-Select Enrichment Principle
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function & Relevance in Clonal Editing Studies |
|---|---|
| CRISPR-Select RNP Complex | Pre-assembled Ribonucleoprotein of Cas9 and target-specific gRNA. Enables rapid, transient editing and is central to the CRISPR-Select protocol. |
| HDR Donor Template with PuroR | A donor DNA construct containing homology arms and the desired edit, linked to a puromycin resistance (PuroR) gene. Allows selection of HDR-successful cells. |
| CloneR or RevitaCell Supplement | Cell culture supplement that enhances single-cell survival post-editing and sorting, critical for clonal outgrowth in all methods. |
| Flow Cytometry Antibodies (e.g., SSEA-4) | For assessing pluripotency marker retention in stem cell models post-editing, a key quality control for clonal lines. |
| NGS-Based Off-Target Kit (e.g., GUIDE-seq) | Comprehensive kit to identify and quantify off-target editing events, essential for validating clonal safety. |
| KaryoStat+ Assay | High-resolution karyotyping service or kit to confirm genomic integrity of expanded clonal lines, especially in therapeutic development. |
Conclusion
The transition from conventional limiting dilution cloning to targeted enrichment methods like CRISPR-Select represents a significant paradigm shift in cell line development. While LDC remains a reliable, instrument-light approach, CRISPR-Select offers a compelling alternative by dramatically accelerating timelines, improving the certainty of monoclonality, and directly linking phenotypic selection to genotypic outcomes. This targeted workflow is particularly transformative for complex editing projects requiring high efficiency, such as generating multiple knockouts or precise knock-ins for therapeutic protein production. Future directions will likely involve the integration of more sophisticated, multi-modal reporters and automated, high-throughput platforms to further enhance scalability. For research and biopharma teams prioritizing speed, precision, and resource efficiency in their gene-editing pipelines, adopting and optimizing CRISPR-Select methodologies will be a critical competitive advantage.